ART-ARH type toxin effector-immunity protein pair of Listeria monocytogenes
Ouyang, S., Hongxin, G., Huan, W.To be published.
Experimental Data Snapshot
Starting Model: in silico
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Lmo0067 protein | 332 | Listeria monocytogenes EGD-e | Mutation(s): 0 Gene Names: lmo0067 | 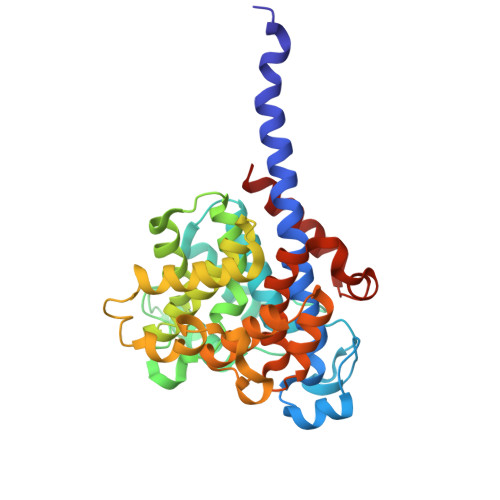 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8YAQ0 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Lmo0066 protein | 180 | Listeria monocytogenes EGD-e | Mutation(s): 0 Gene Names: lmo0066 | 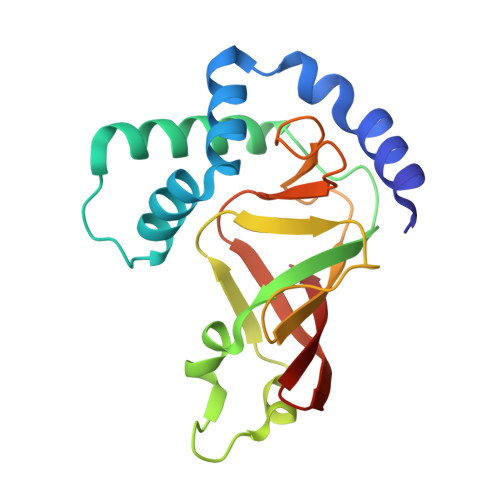 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8YAQ1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| AR6 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth A] | [(2R,3S,4R,5R)-5-(6-AMINOPURIN-9-YL)-3,4-DIHYDROXY-OXOLAN-2-YL]METHYL[HYDROXY-[[(2R,3S,4R,5S)-3,4,5-TRIHYDROXYOXOLAN-2-YL]METHOXY]PHOSPHORYL] HYDROGEN PHOSPHATE C15 H23 N5 O14 P2 SRNWOUGRCWSEMX-ZQSHOCFMSA-N |  | ||
| MG (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A], E [auth A] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 84.02 | α = 90 |
| b = 84.02 | β = 90 |
| c = 82.81 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-3000 | data reduction |
| HKL-3000 | data scaling |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | 31271265 |