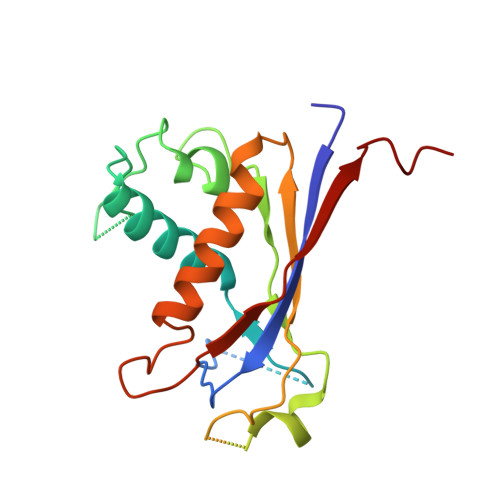
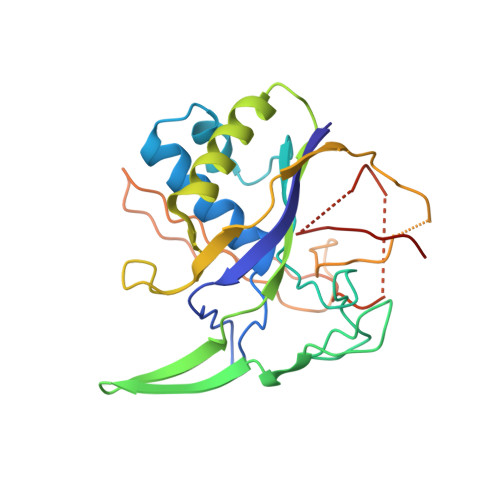
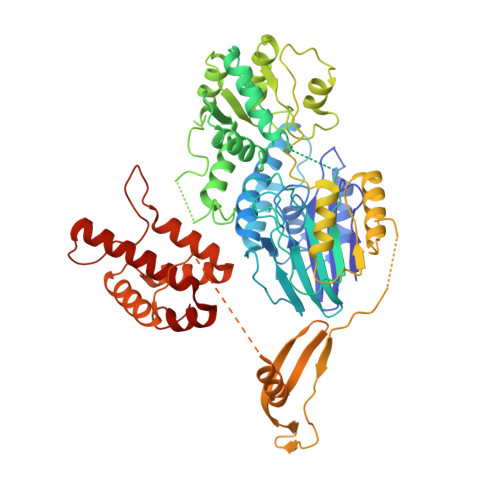
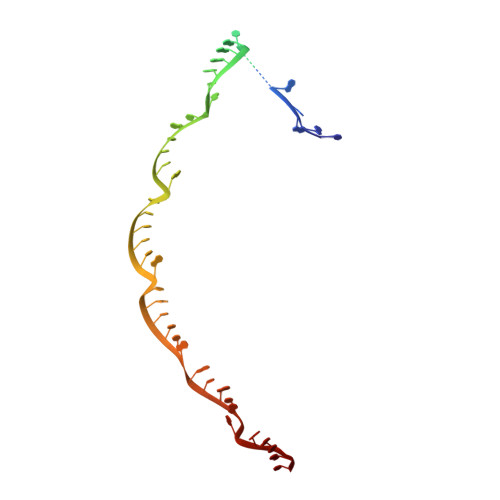
Structural basis for the activity of the type VII CRISPR-Cas system.
Yang, J., Li, X., He, Q., Wang, X., Tang, J., Wang, T., Zhang, Y., Yu, F., Zhang, S., Liu, Z., Zhang, L., Liao, F., Yin, H., Zhao, H., Deng, Z., Zhang, H.(2024) Nature 633: 465-472
- PubMed: 39143216 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-024-07815-0
- Primary Citation Related Structures:
8YHD, 8YHE, 8Z4J, 8Z4L, 8Z99, 8Z9C, 8Z9E - PubMed Abstract:
The newly identified type VII CRISPR-Cas candidate system uses a CRISPR RNA-guided ribonucleoprotein complex formed by Cas5 and Cas7 proteins to target RNA 1 . However, the RNA cleavage is executed by a dedicated Cas14 nuclease, which is distinct from the effector nucleases of the other CRISPR-Cas systems. Here we report seven cryo-electron microscopy structures of the Cas14-bound interference complex at different functional states. Cas14, a tetrameric protein in solution, is recruited to the Cas5-Cas7 complex in a target RNA-dependent manner. The N-terminal catalytic domain of Cas14 binds a stretch of the substrate RNA for cleavage, whereas the C-terminal domain is primarily responsible for tethering Cas14 to the Cas5-Cas7 complex. The biochemical cleavage assays corroborate the captured functional conformations, revealing that Cas14 binds to different sites on the Cas5-Cas7 complex to execute individual cleavage events. Notably, a plugged-in arginine of Cas7 sandwiched by a C-shaped clamp of C-terminal domain precisely modulates Cas14 binding. More interestingly, target RNA cleavage is altered by a complementary protospacer flanking sequence at the 5' end, but not at the 3' end. Altogether, our study elucidates critical molecular details underlying the assembly of the interference complex and substrate cleavage in the type VII CRISPR-Cas system, which may help rational engineering of the type VII CRISPR-Cas system for biotechnological applications.
- State Key Laboratory of Experimental Hematology, Key Laboratory of Immune Microenvironment and Disease (Ministry of Education), International Joint Laboratory of Ocular Diseases (Ministry of Education), The Province and Ministry Co-sponsored Collaborative Innovation Center for Medical Epigenetics, Tianjin Institute of Immunology, School of Basic Medical Sciences, Tianjin Medical University, Tianjin, China.
Organizational Affiliation: