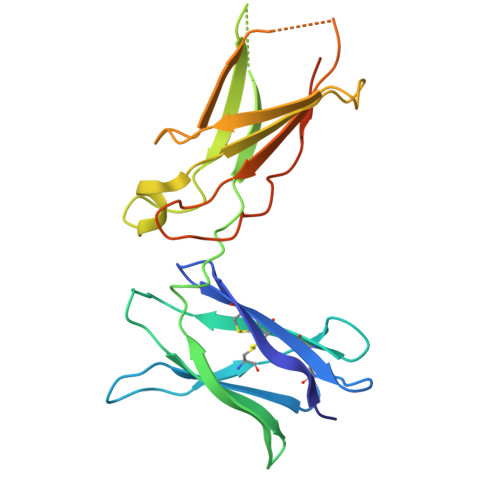
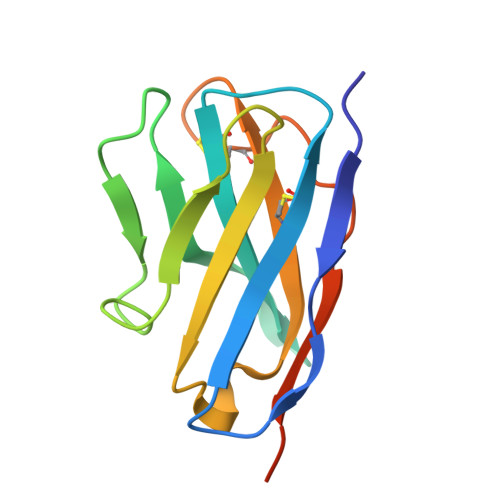
Structural insight into interleukin-4R alpha and interleukin-5 inhibition by nanobodies from a bispecific antibody.
Qiu, W., Meng, J., Su, Z., Xie, W., Song, G.(2024) MedComm (2020) 5: e700-e700
- PubMed: 39252822 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/mco2.700
- Primary Citation Related Structures:
8Z8L - Shanghai Key Laboratory of Regulatory Biology Institute of Biomedical Sciences and School of Life Sciences East China Normal University Shanghai China.
Organizational Affiliation: