Structural insights into the activation mechanism of the human metabolite receptor HCAR1.
Gao, M., Zang, S., Zhu, Y., Xi, K., Du, Y., Cheng, S., Miao, L., Lu, Y., Mao, C., Zhang, Y., Ma, X.(2026) Sci Signal 19: eadw1483-eadw1483
- PubMed: 41493973 Search on PubMed
- DOI: https://doi.org/10.1126/scisignal.adw1483
- Primary Citation Related Structures:
8Z87, 8Z8A, 8Z8B - PubMed Abstract:
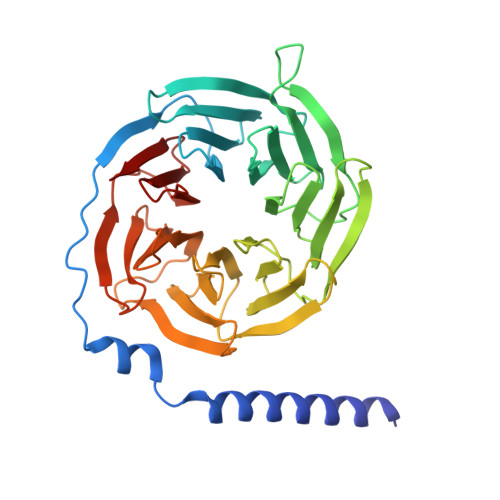
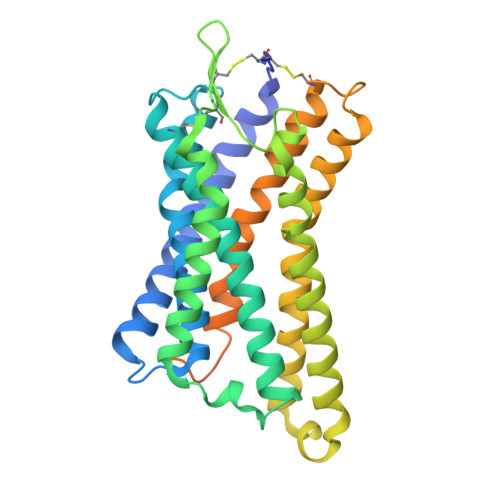
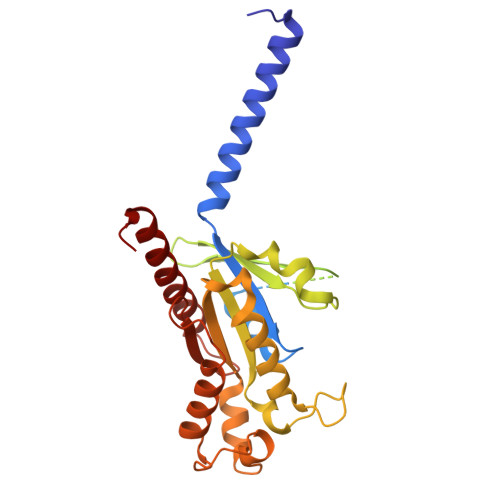
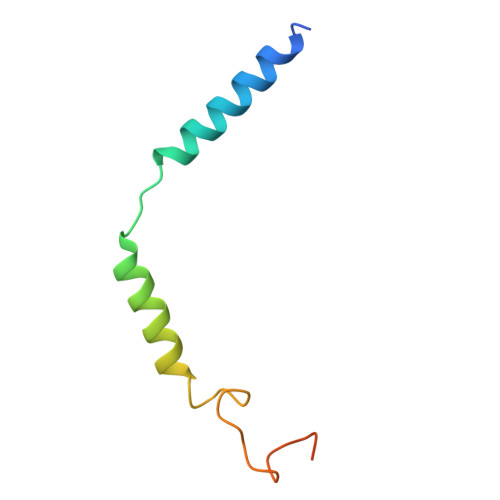
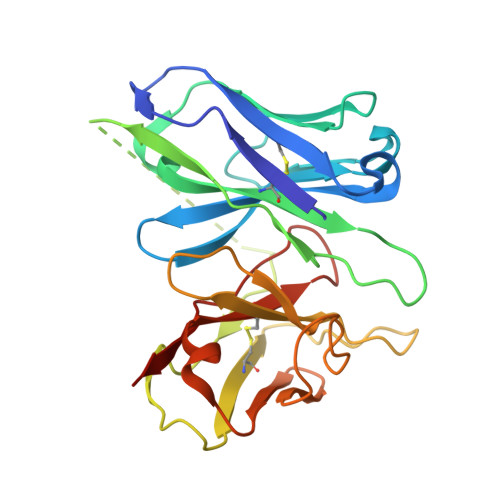
Hydroxycarboxylic acid receptor 1 (HCAR1) is a class A G protein-coupled receptor (GPCR) that is activated by the endogenous metabolite l-lactate and that plays an important role in various metabolic and inflammatory disorders. HCAR1 uses distinct ligand recognition and self-activation mechanisms to mediate specific pathophysiological functions through Gα i/o and β-arrestin signaling pathways. To support effective drug development targeting HCAR1, we investigated ligand recognition and activation mechanisms through cryo-electron microscopy (cryo-EM) structures of the HCAR1-Gα i1 complex in the apo state or with l-lactate or with the synthetic agonist CHBA. Compared with other HCARs, HCAR1 has a more compact binding pocket, which is stabilized by three unique disulfide bonds. l-lactate exhibited a flexible binding mode and relatively weak intermolecular interactions, thus requiring millimolar concentrations for receptor activation. In contrast, the binding of CHBA was more stable because of its chlorinated benzene ring, thus resulting in improved agonist potency. Structural comparisons with HCAR2 identified critical residues that restrict the size of the binding pocket of HCAR1 and influence ligand selectivity. Self-activation of HCAR1 is driven by conformational rearrangements within extracellular loop 2, with Phe168 ECL2 playing a pivotal role as the key agonist. Together, these results clarify the mechanisms underlying HCAR1 activation, self-activation, and ligand selectivity, providing a structural framework for the design of high-affinity, selective agonists and inverse agonists with minimized off-target effects.
- MOE Medical Basic Research Innovation Center for Gut Microbiota and Chronic Diseases, School of Medicine, Jiangnan University, Wuxi 214122, China.
Organizational Affiliation: