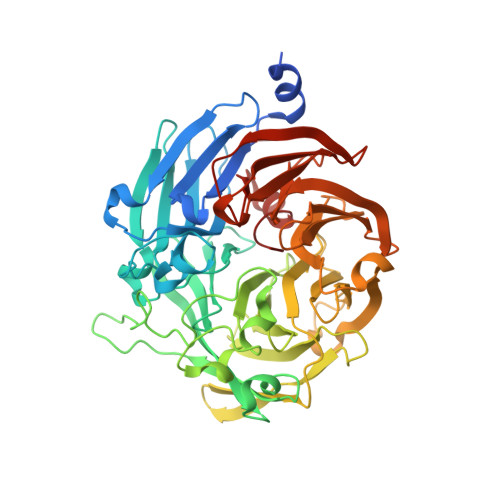
Effect of Copper Ions on the Crystal Packing and Conformation of Thiocyanate Dehydrogenase in the Crystal Structure
Varfolomeeva, L.A., Solovieva, A.Y., Shipkov, N.S., Dergousova, N.I., Minyaev, M.E., Boyko, K.M., Tikhonova, T.V., Popov, V.O.(2025) Crystallogr Rep 70: 8-15