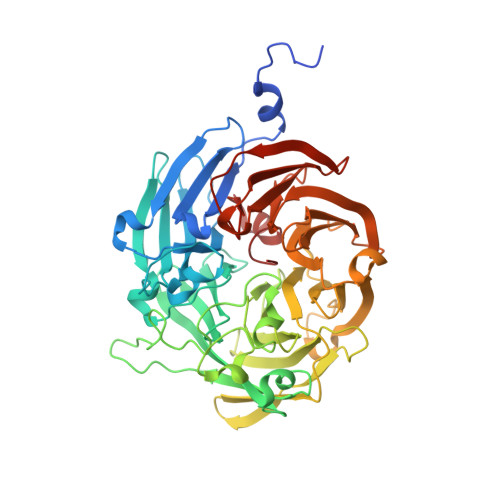
The structure of non-activated thiocyanate dehydrogenase mutant with the H447Q substitution from Pelomicrobium methylotrophicum (pmTcDH H447Q)
Varfolomeeva, L.A., Shipkov, N.S., Dergousova, N.I., Boyko, K.M., Tikhonova, T.V., Popov, V.O.To be published.