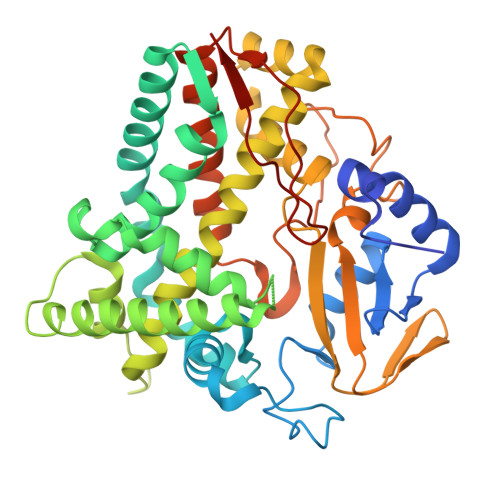
Structural Insights into the Interaction of Terpenoids with Streptomyces avermitilis CYP107P2.
Jeong, E., Kim, V., Kim, C., Lee, Y.B., Kim, D.(2024) Biomol Ther (Seoul) 32: 474-480
- PubMed: 38835149 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.4062/biomolther.2024.045
- Primary Citation Related Structures:
8YHN - PubMed Abstract:
Streptomyces avermitilis genome includes 33 genes encoding monooxygenation-catalyzing cytochrome P450 enzymes. We investigated the structure of CYP107P2 and its interactions with terpenoid compounds. The recombinant CYP107P2 protein was expressed in Escherichia coli and the purified enzyme exhibited a typical P450 spectrum upon CO-binding in its reduced state. Type-I substrate-binding spectral titrations were observed with various terpenoid compounds, including α-pinene, β-pinene, α-terpinyl acetate, and (+)-3-carene. The calculated binding affinities ( K d ) ranged from 15.9 to 50.8 μM. The X-ray crystal structure of CYP107P2 was determined at 1.99 Å resolution, with a well-conserved overall P450 folding conformation. The terpenoid compound docking models illustrated that the structural interaction between monoterpenes and CYP107P2, with the distance between heme and terpenes ranging from 3.4 to 5.4 Å, indicates potential substrate binding for P450 enzyme. This study suggests that CYP107P2 is a Streptomyces P450 enzyme capable of catalyzing terpenes as substrates, signifying noteworthy advancements in comprehending a novel P450 enzyme's involvement in terpene reactions.
- Department of Biological Sciences, Konkuk University, Seoul 05025, Republic of Korea.
Organizational Affiliation: