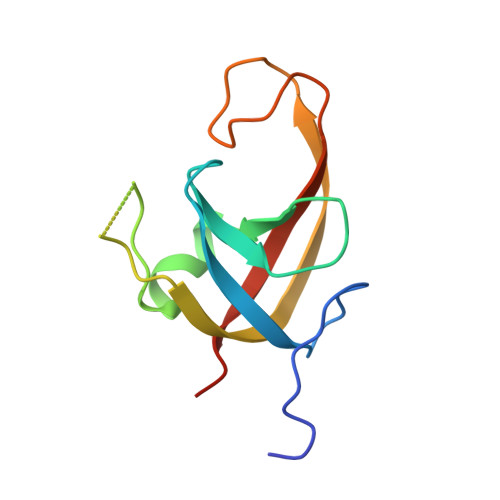
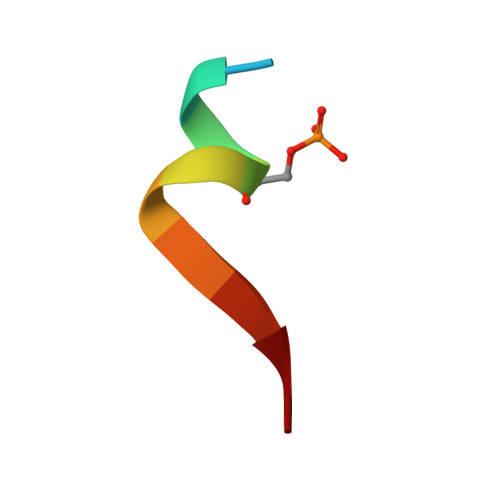Structural basis for TNIP1 binding to FIP200 during mitophagy.
Wu, S., Li, M., Wang, L., Yang, L., Cui, J., Li, F., Wang, Q., Shi, Y., Lv, M.(2024) J Biological Chem 300: 107605-107605
- PubMed: 39059492 Search on PubMed
- DOI: https://doi.org/10.1016/j.jbc.2024.107605
- Primary Citation Related Structures:
8YFK, 8YFL, 8YFM, 8YFN - PubMed Abstract:
TNIP1 has been increasingly recognized as a security check to finely adjust the rate of mitophagy by disrupting the recycling of the Unc-51-like kinase (ULK) complex during autophagosome formation. Through tank-binding kinase 1 (TBK1)-mediated phosphorylation of the TNIP1 FIR motif, the binding affinity of TNIP1 for FIP200, a component of the ULK complex, is enhanced, allowing TNIP1 to outcompete autophagy receptors. Consequently, FIP200 is released from the autophagosome, facilitating further autophagosome expansion. However, the molecular basis by which FIP200 utilizes its claw domain to distinguish the phosphorylation status of residues in the TNIP1 FIP200 interacting region (FIR) motif for recognition is not well understood. Here, we elucidated multiple crystal structures of the complex formed by the FIP200 claw domain and various phosphorylated TNIP1 FIR peptides. Structural and isothermal titration calorimetry (ITC) analyses identified the crucial residues in the FIP200 claw domain responsible for the specific recognition of phosphorylated TNIP1 FIR peptides. Additionally, utilizing structural comparison and molecular dynamics (MD) simulation data, we demonstrated that the C-terminal tail of TNIP1 peptide affected its binding to the FIP200 claw domain. Moreover, the phosphorylation of TNIP1 Ser123 enabled the peptide to effectively compete with the peptide p-CCPG1 (the FIR motif of the autophagy receptor CCPG1) for binding with the FIP200 claw domain. Overall, our work provides a comprehensive understanding of the specific recognition of phosphorylated TNIP1 by the FIP200 claw domain, marking an initial step toward fully understanding the molecular mechanism underlying the TNIP1-dependent inhibition of mitophagy.
- Hefei National Research Center for Cross Disciplinary Science, School of Life Sciences, Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei, Anhui 230026, China; Ministry of Education Key Laboratory for Membraneless Organelles and Cellular Dynamics, University of Science & Technology of China, Hefei, Anhui 230026, China; Center for Advanced Interdisciplinary Science and Biomedicine of IHM, Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei, Anhui 230026, China.
Organizational Affiliation: