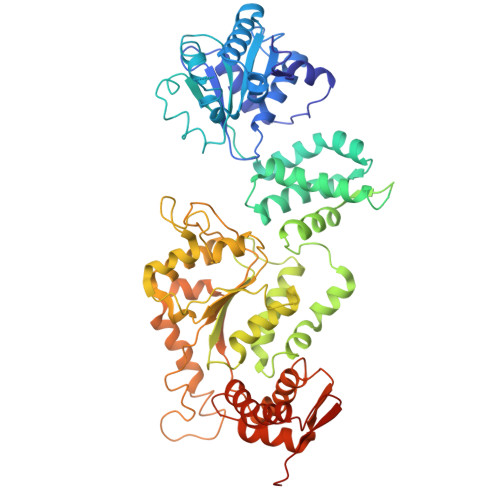
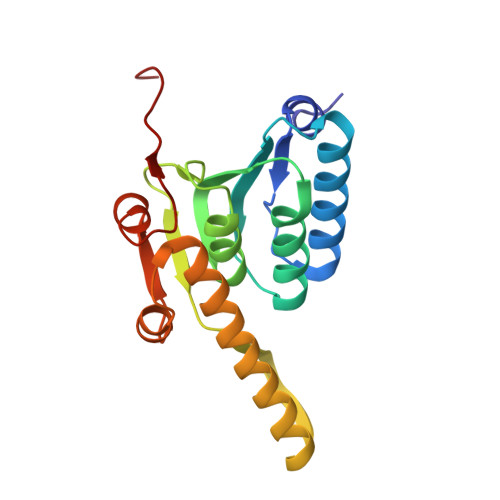
Structural Insights into Bortezomib-Induced Activation of the Caseinolytic Chaperone-Protease System in Mycobacterium tuberculosis
Zhou, B., Gao, Y., Zhao, H., Liu, B., Zhang, H., Fang, C., Yuan, H., Wang, J., Li, Z., Zhao, Y., Huang, X., Wang, X., Oliveira, A.S.F., Spencer, J., Mulholland, A.J., Burston, S.G., Hu, J., Su, N., Chen, X., He, J., Zhang, T., Xiong, X.(2025) Nat Commun 16: 3466