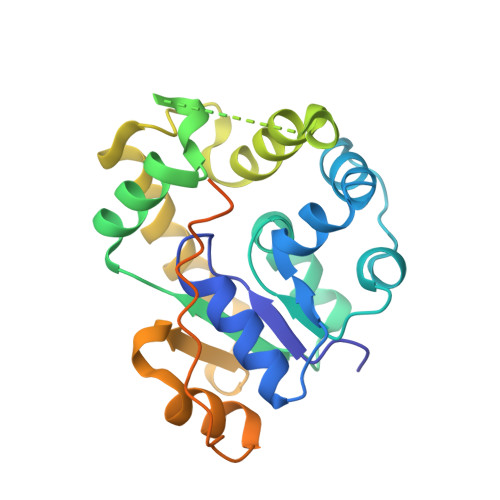
Crystal structure of activating sulfotransferase SgdX2 involved in biosynthesis of secondary metabolite sungeidine.
Mori, T., Teramoto, T., Kakuta, Y.(2024) Biochem Biophys Res Commun 711: 149891-149891
- PubMed: 38621346 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2024.149891
- Primary Citation Related Structures:
8Y9P - PubMed Abstract:
Microorganisms synthesize a plethora of complex secondary metabolites, many of which are beneficial to human health, such as anticancer agents and antibiotics. Among these, the Sungeidines are a distinct class of secondary metabolites known for their bulky and intricate structures. They are produced by a specific biosynthetic gene cluster within the genome of the soil-dwelling actinomycete Micromonospora sp. MD118. A notable enzyme in the Sungeidine biosynthetic pathway is the activating sulfotransferase SgdX2. In this pathway, SgdX2 mediates a key sulfation step, after which the product undergoes spontaneous dehydration to yield a Sungeidine compound. To delineate the structural basis for SgdX2's substrate recognition and catalytic action, we have determined the crystal structure of SgdX2 in complex with its sulfate donor product, 3'-phosphoadenosine 5'-phosphate (PAP), at a resolution of 1.6 Å. Although SgdX2 presents a compact overall structure, its core elements are conserved among other activating sulfotransferases. Our structural analysis reveals a unique substrate-binding pocket that accommodates bulky, complex substrates, suggesting a specialized adaptation for Sungeidine synthesis. Moreover, we have constructed a substrate docking model that provides insights into the molecular interactions between SgdX2 and Sungeidine F, enhancing our understanding of the enzyme's specificity and catalytic mechanism. The model supports a general acid-base catalysis mechanism, akin to other sulfotransferases, and underscores the minor role of disordered regions in substrate recognition. This integrative study of crystallography and computational modeling advances our knowledge of microbial secondary metabolite biosynthesis and may facilitate the development of novel biotechnological applications.
- Laboratory of Biophysical Chemistry, Department of Bioscience and Biotechnology, Faculty of Agriculture, Kyushu University, 744 Motooka, Nishi-ku, Fukuoka, 819-0395, Japan.
Organizational Affiliation: