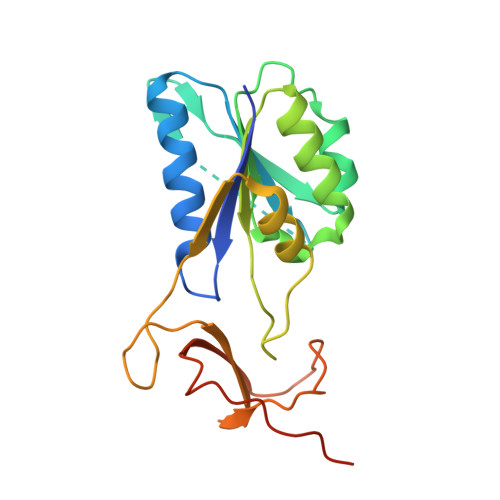
Crystal structure of thymidine kinase from the multi-drug resistant col strain of Staphylococcus aureus.
Ashraf, A., Pal, R.K., Hassan, M.I.(2025) Biochim Biophys Acta Proteins Proteom 1873: 141071-141071
- PubMed: 40189173 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2025.141071
- Primary Citation Related Structures:
8Y7W - PubMed Abstract:
Thymidine kinase (TK) is a key enzyme in the salvage pathway of thymidine that produces thymidine monophosphate. TK enzyme activity is tightly coupled to the cell cycle, exhibiting marked fluctuations in expression and activity. We report the crystal structure of TK from the Staphylococcus aureus col strain (Sa-TK), which has emerged as a promising therapeutic target. The overall structure of Sa-TK closely resembles that of human TK. The lasso region in the structure shows an open conformation due to the absence of a natural substrate. The phosphate donor site is bound with sulfate ions from the crystallization conditions. The P-loop is visible, but the complete P-β hairpin cannot be traced due to the flexibility of this region. Sa-TK assembles as a tetramer with unique inter-subunit interactions involving salt bridges between charged residues. Glu136 and Arg184, as well as Arg154 and Glu102 from each of the subunits, have β-sheet interactions that form salt bridges. The catalytically active site residue Glu89 is conserved, which is essential for enzyme activity. Sa-TK lacks a longer C-terminal sequence involved in mitotic regulation through proteolytic degradation, a feature that is likely absent in Sa-TK. The crystal structure of Sa-TK offers detailed insights into its structural and functional properties, highlighting its conserved nature and emphasizing the challenge of developing selective inhibitors that do not affect host TK. This detailed structural information presents a valuable opportunity for the rational design of novel antibacterial agents specifically targeting Sa-TK, offering a promising avenue for combating S. aureus infections.
- Centre for Interdisciplinary Research in Basic Sciences, Jamia Millia Islamia, Jamia Nagar, New Delhi 110025, India. Electronic address: ashraf.anam6@gmail.com.
Organizational Affiliation: