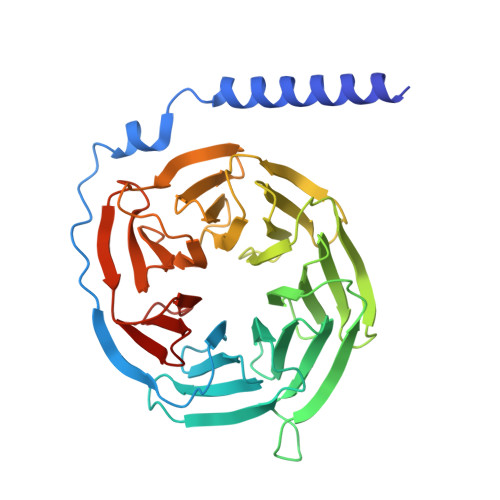
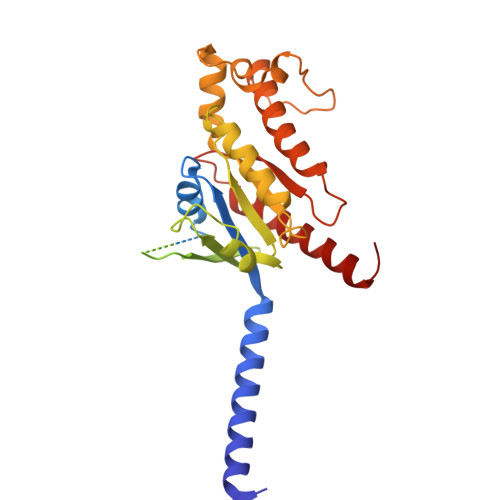
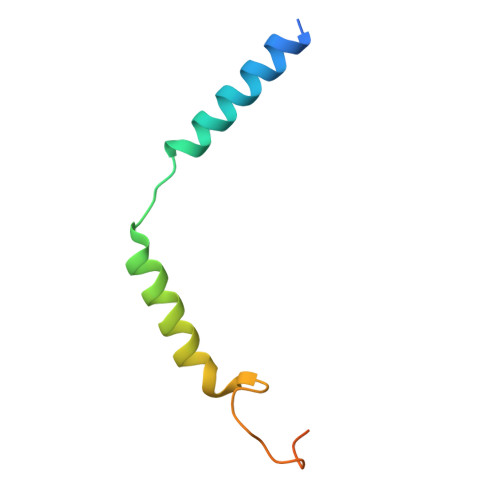
Structural insights into ligand recognition, selectivity, and activation of bombesin receptor subtype-3.
Li, C., Xu, Y., Su, W., He, X., Li, J., Li, X., Xu, H.E., Yin, W.(2024) Cell Rep 43: 114511-114511
- PubMed: 39024101 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2024.114511
- Primary Citation Related Structures:
8Y52, 8Y53 - PubMed Abstract:
Bombesin receptor subtype-3 (BRS3) is an important orphan G protein-coupled receptor that regulates energy homeostasis and insulin secretion. As a member of the bombesin receptor (BnR) family, the lack of known endogenous ligands and high-resolution structure has hindered the understanding of BRS3 signaling and function. We present two cryogenic electron microscopy (cryo-EM) structures of BRS3 in complex with the heterotrimeric G q protein in its active states: one bound to the pan-BnR agonist BA1 and the other bound to the synthetic BRS3-specific agonist MK-5046. These structures reveal the architecture of the orthosteric ligand pocket underpinning molecular recognition and provide insights into the structural basis for BRS3's selectivity and low affinity for bombesin peptides. Examination of conserved micro-switches suggests a shared activation mechanism among BnRs. Our findings shed light on BRS3's ligand selectivity and signaling mechanisms, paving the way for exploring its therapeutic potential for diabetes, obesity, and related metabolic disorders.
- State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai 201203, China; School of Life Science and Technology, ShanghaiTech University, Shanghai 201210, China; Lingang Laboratory, Shanghai 200031, China.
Organizational Affiliation: