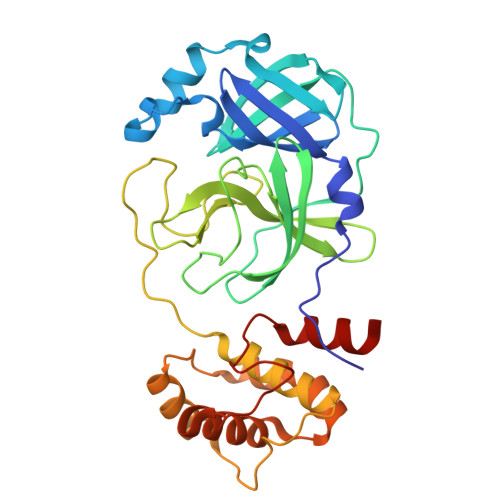
Discovery of Novel Nonpeptidic and Noncovalent Small Molecule 3CL pro Inhibitors as anti-SARS-CoV-2 Drug Candidate.
Jiang, Z., Feng, B., Chen, L., Nie, T., Chen, S., Wang, L., Liu, H., Yu, T., Zhang, Y., Zheng, M., Xu, Y., Liu, H., Zang, Y., Su, H., Zhang, L., Li, J., Zhou, Y.(2024) J Med Chem 67: 12760-12783
- PubMed: 39072488 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.4c00739
- Primary Citation Related Structures:
8GTV, 8GTW, 8Y42, 8Y44 - PubMed Abstract:
SARS-CoV-2 has still been threatening global public health with its emerging variants. Our previous work reported lead compound JZD-07 that displayed good 3CL pro inhibitory activity. Here, an in-depth structural optimization for JZD-07 was launched to obtain more desirable drug candidates for the therapy of SARS-CoV-2 infection, in which 54 novel derivatives were designed and synthesized by a structure-based drug design strategy. Among them, 24 compounds show significantly enhanced 3CL pro inhibitory potencies with IC 50 values less than 100 nM, and 11 compounds dose-dependently inhibit the replication of the SARS-CoV-2 delta variant. In particular, compound 49 has the most desirable antiviral activity with EC 50 of 0.272 ± 0.013 μM against the delta variant, which was more than 20 times stronger than JZD-07 . Oral administration of 49 could significantly reduce the lung viral copies of mice, exhibiting a more favorable therapeutic potential. Overall, this investigation presents a promising drug candidate for further development to treat SARS-CoV-2 infection.
- Lingang Laboratory, Shanghai 200031, China.
Organizational Affiliation: