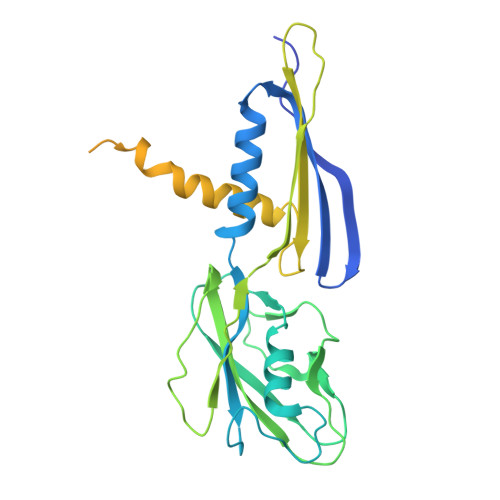
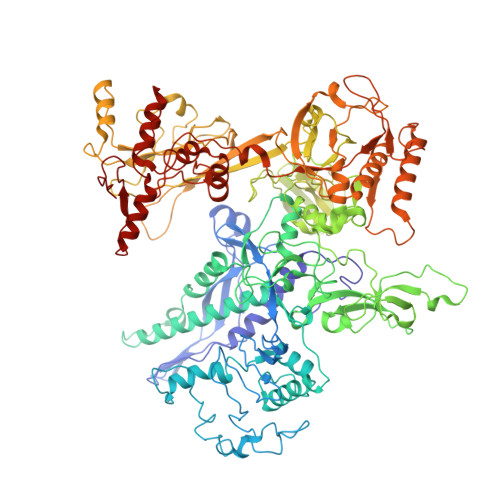
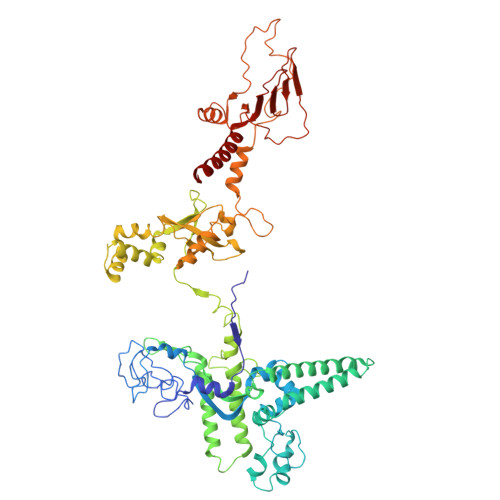
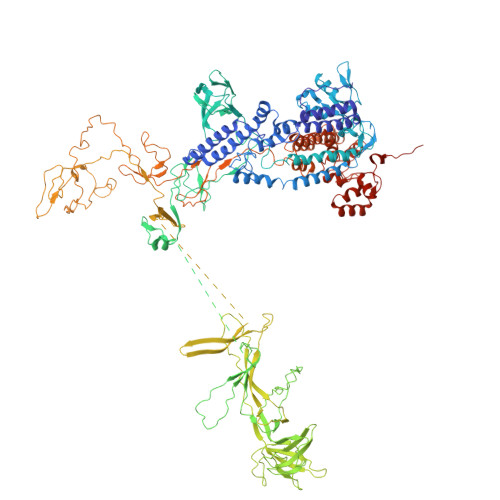
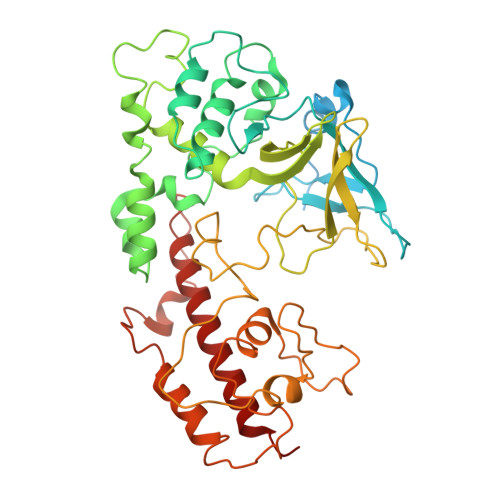
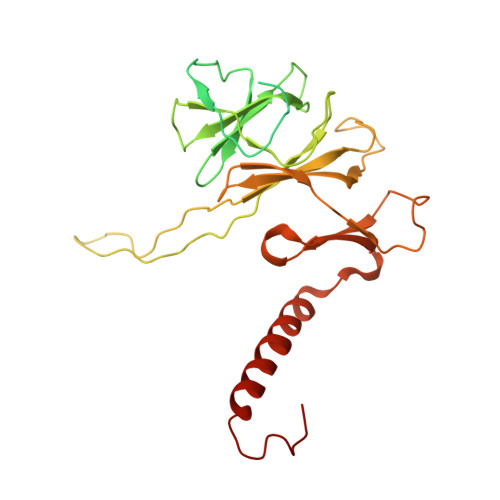
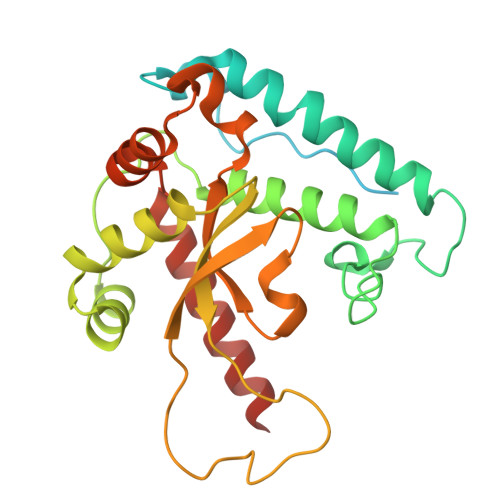
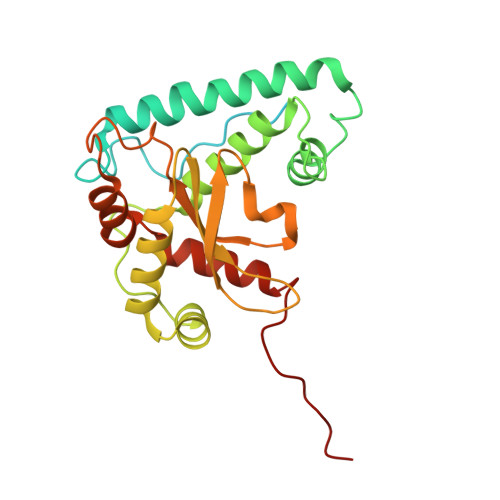
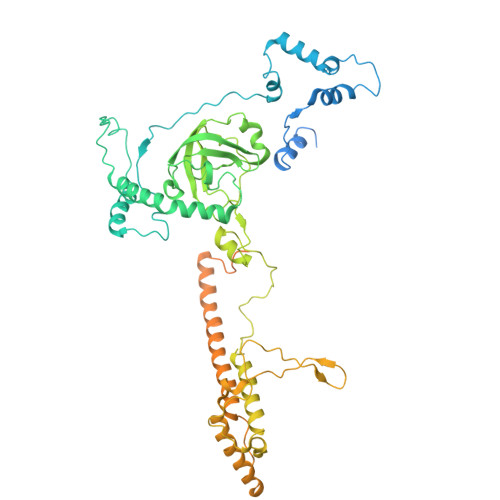
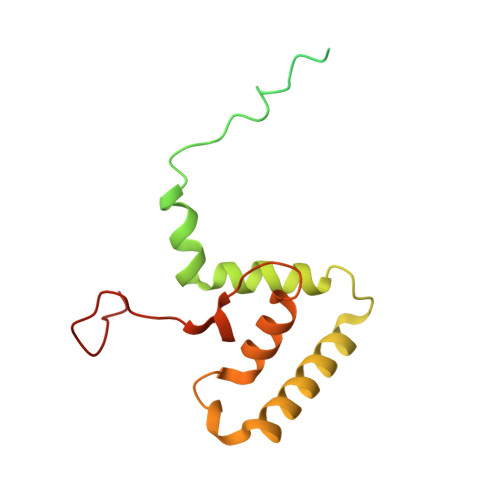
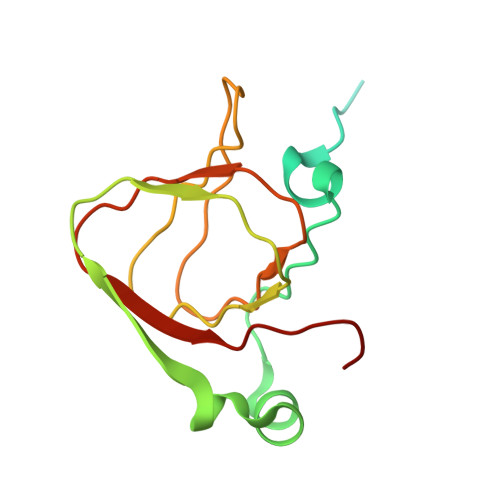
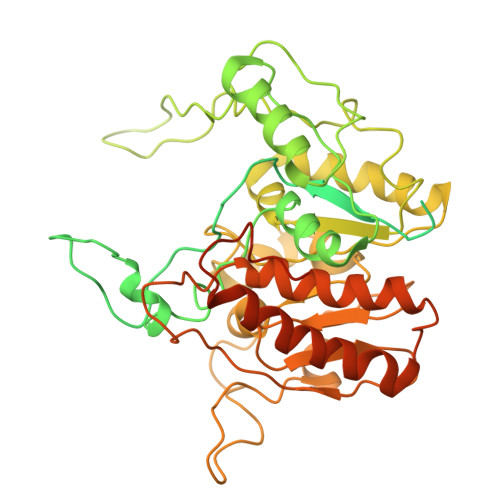
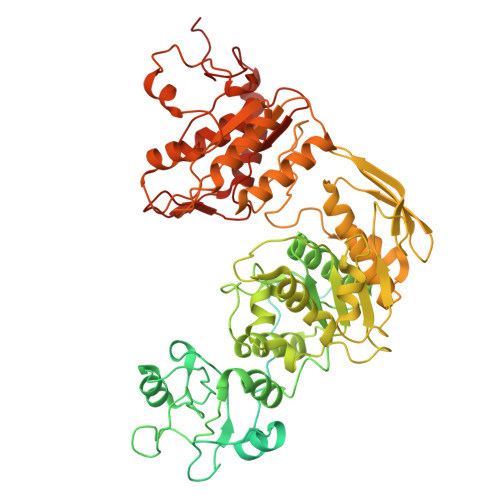
Architecture of the spinach plastid-encoded RNA polymerase.
Wang, T., Wang, G.L., Fang, Y., Zhang, Y., Peng, W., Zhou, Y., Zhang, A., Yu, L.J., Lu, C.(2024) Nat Commun 15: 9838-9838
- PubMed: 39537621 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-54266-2
- Primary Citation Related Structures:
8XZV - PubMed Abstract:
The plastid-encoded RNA polymerase serves as the principal transcription machinery within chloroplasts, transcribing over 80% of all primary plastid transcripts. This polymerase consists of a prokaryotic-like core enzyme known as the plastid-encoded RNA polymerase core, and is supplemented by newly evolved associated proteins known as PAPs. However, the architecture of the plastid-encoded RNA polymerase and the possible functions of PAPs remain unknown. Here, we present the cryo-electron microscopy structure of a 19-subunit plastid-encoded RNA polymerase complex derived from spinach (Spinacia oleracea). The structure shows that the plastid-encoded RNA polymerase core resembles bacterial RNA polymerase. Twelve PAPs and two additional proteins (FLN2 and pTAC18) bind at the periphery of the plastid-encoded RNA polymerase core, forming extensive interactions that may facilitate complex assembly and stability. PAPs may also protect the complex against oxidative damage and has potential functions in transcriptional regulation. This research offers a structural basis for future investigations into the functions and regulatory mechanisms governing the transcription of plastid genes.
- State Key Laboratory of Wheat Improvement, College of Life Sciences, Shandong Agricultural University, Taian, Shandong, 271018, China.
Organizational Affiliation: