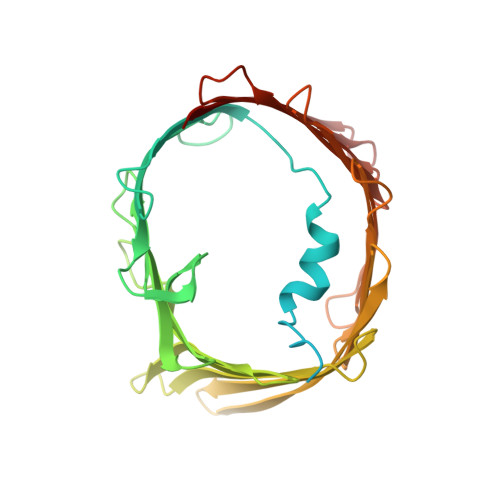
Structure of the intact Tom20 receptor in the human translocase of the outer membrane complex.
Su, J., Tian, X., Wang, Z., Yang, J., Sun, S., Sui, S.F.(2024) PNAS Nexus 3: pgae269-pgae269
- PubMed: 39071881 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/pnasnexus/pgae269
- Primary Citation Related Structures:
8XVA - PubMed Abstract:
The translocase of the outer membrane (TOM) complex serves as the main gate for preproteins entering mitochondria and thus plays a pivotal role in sustaining mitochondrial stability. Precursor proteins, featuring amino-terminal targeting signals (presequences) or internal targeting signals, are recognized by the TOM complex receptors Tom20, Tom22, and Tom70, and then translocated into mitochondria through Tom40. By using chemical cross-linking to stabilize Tom20 in the TOM complex, this study unveils the structure of the human TOM holo complex, encompassing the intact Tom20 component, at a resolution of approximately 6 Å by cryo-electron microscopy. Our structure shows the TOM holo complex containing only one Tom20 subunit, which is located right at the center of the complex and stabilized by extensive interactions with Tom22, Tom40, and Tom6. Based on the structure, we proposed a possible translocation mode of TOM complex, by which different receptors could work simultaneously to ensure that the preproteins recognized by them are all efficiently translocated into the mitochondria.
- State Key Laboratory of Membrane Biology, Beijing Frontier Research Center for Biological Structure, Beijing Advanced Innovation Center for Structural Biology, School of Life Sciences, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: