Activation of DNMT1 by Chemically Synthesized Dual monoubiquitinated PAF15 Protein
Hayashi, G., Takahashi, Y., Kikuchi, A., Kori, S., Arita, K., Murakami, H.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA (cytosine-5)-methyltransferase 1 | 1,271 | Homo sapiens | Mutation(s): 0 Gene Names: DNMT1, AIM, CXXC9, DNMT EC: 2.1.1.37 | 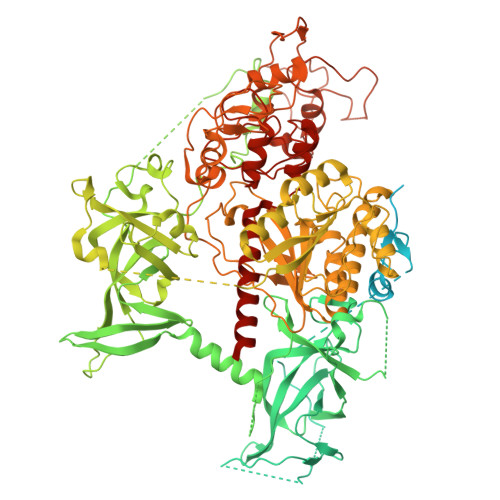 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P26358 GTEx: ENSG00000130816 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P26358 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(*AP*CP*TP*TP*AP*(5CM)P*GP*GP*AP*AP*GP*G)-3') | 12 | synthetic construct | 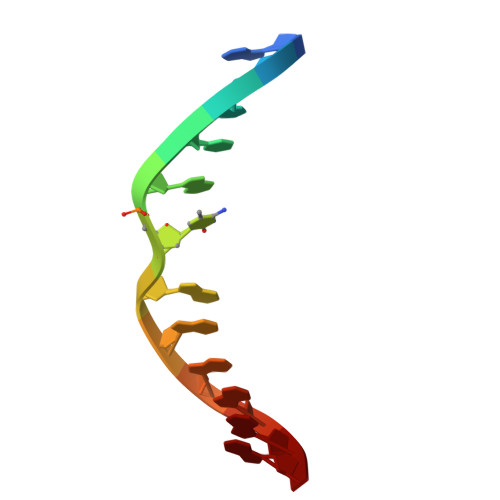 | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 3 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(*CP*CP*TP*TP*CP*(C55)P*GP*TP*AP*AP*GP*T)-3') | 12 | synthetic construct | 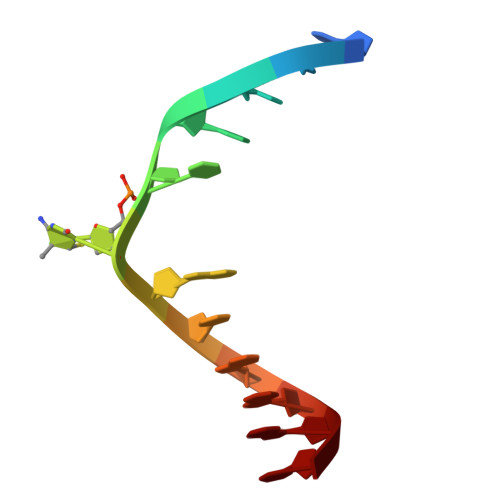 | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SAH (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A] | S-ADENOSYL-L-HOMOCYSTEINE C14 H20 N6 O5 S ZJUKTBDSGOFHSH-WFMPWKQPSA-N |  | ||
| ZN (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | E [auth A], F [auth A] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Japan Society for the Promotion of Science (JSPS) | Japan | 18H02392, 19H05294, 19H05741,19J22030 |