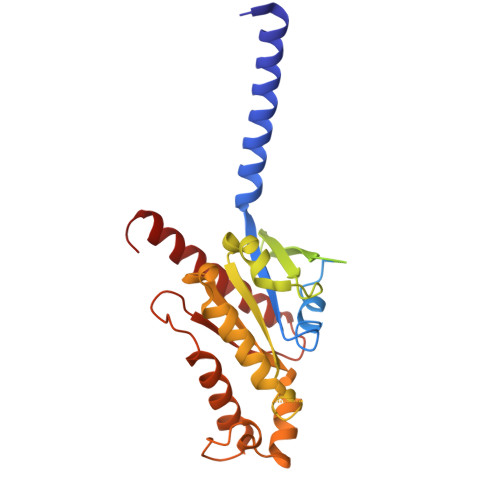
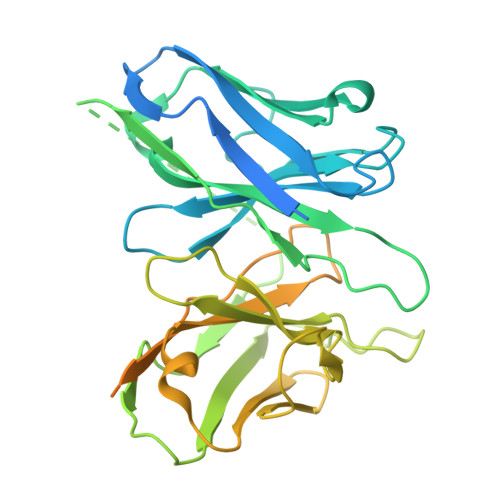
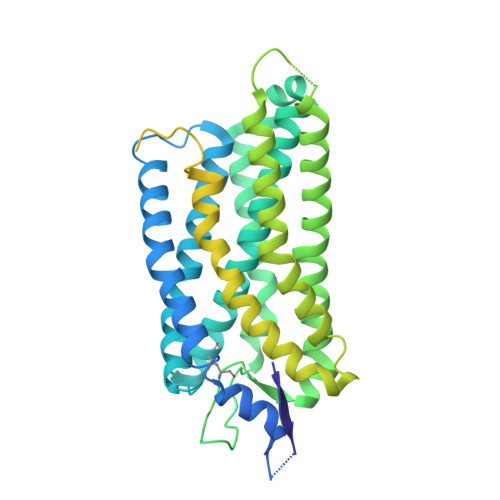
Structural basis of tethered agonism and G protein coupling of protease-activated receptors.
Guo, J., Zhou, Y.L., Yang, Y., Guo, S., You, E., Xie, X., Jiang, Y., Mao, C., Xu, H.E., Zhang, Y.(2024) Cell Res 34: 725-734
- PubMed: 38997424 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-024-00997-2
- Primary Citation Related Structures:
8XOR, 8XOS - PubMed Abstract:
Protease-activated receptors (PARs) are a unique group within the G protein-coupled receptor superfamily, orchestrating cellular responses to extracellular proteases via enzymatic cleavage, which triggers intracellular signaling pathways. Protease-activated receptor 1 (PAR1) is a key member of this family and is recognized as a critical pharmacological target for managing thrombotic disorders. In this study, we present cryo-electron microscopy structures of PAR1 in its activated state, induced by its natural tethered agonist (TA), in complex with two distinct downstream proteins, the G q and G i heterotrimers, respectively. The TA peptide is positioned within a surface pocket, prompting PAR1 activation through notable conformational shifts. Contrary to the typical receptor activation that involves the outward movement of transmembrane helix 6 (TM6), PAR1 activation is characterized by the simultaneous downward shift of TM6 and TM7, coupled with the rotation of a group of aromatic residues. This results in the displacement of an intracellular anion, creating space for downstream G protein binding. Our findings delineate the TA recognition pattern and highlight a distinct role of the second extracellular loop in forming β-sheets with TA within the PAR family, a feature not observed in other TA-activated receptors. Moreover, the nuanced differences in the interactions between intracellular loops 2/3 and the Gα subunit of different G proteins are crucial for determining the specificity of G protein coupling. These insights contribute to our understanding of the ligand binding and activation mechanisms of PARs, illuminating the basis for PAR1's versatility in G protein coupling.
- Department of Pharmacology and Department of Pathology of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine, Hangzhou, Zhejiang, China.
Organizational Affiliation: