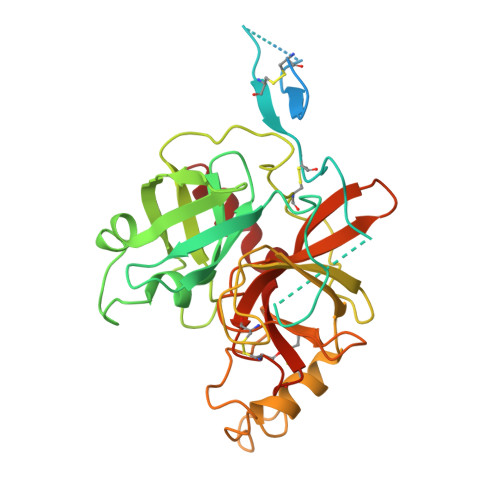
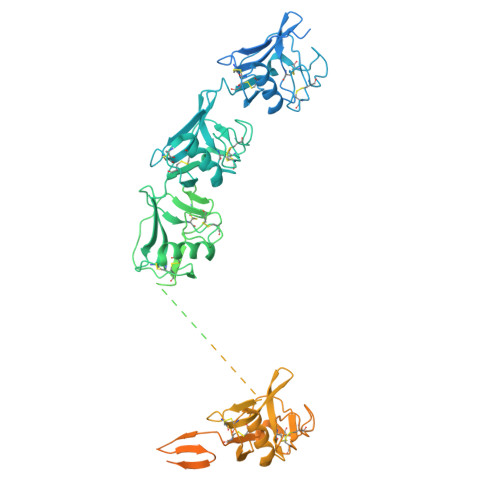
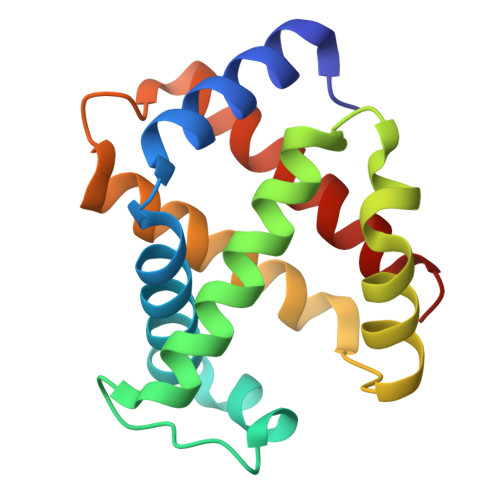
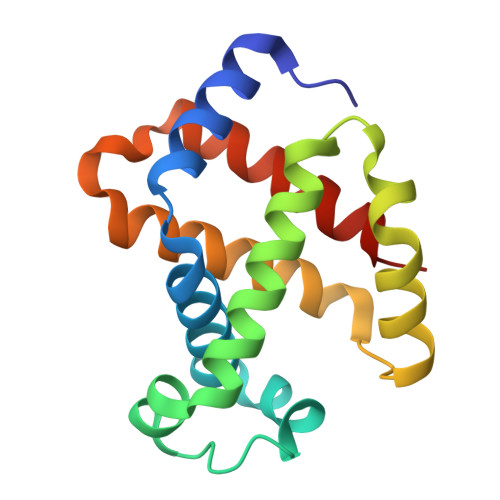
Calcium-dependent oligomerization of scavenger receptor CD163 facilitates the endocytosis of ligands.
Xu, H., Song, X., Su, X.D.(2025) Nat Commun 16: 6679-6679
- PubMed: 40691148 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-62013-4
- Primary Citation Related Structures:
8XMK, 8XMP, 8XMQ, 8XMW - PubMed Abstract:
Scavenger receptor CD163 is a marker of M2 type macrophages that play important roles in anti-inflammatory processes. The most extensively studied function of CD163 is related to the elimination of hemoglobin-haptoglobin (Hb-Hp) complexes, to prevent potential oxidative toxicity of the iron-containing heme. However, the structural mechanism of CD163 in ligand binding and internalization remains elusive. Here, we present the cryo-electron microscopy structure of human Hb-Hp recognition by the full ectodomain of CD163. We illuminate that CD163 forms calcium-dependent oligomers and primarily exists as trimeric form under the condition of 2.5 mM calcium. It mainly utilizes two protomers to interact with Hb-Hp complex asymmetrically, while the third protomer of the trimer also has the potential to form calcium-mediated contacts with Hp. Flow cytometry analyses reveal that oligomerization of CD163 significantly enhances the efficiency of ligand endocytosis. These results advance our understanding of the role of CD163 in ligand scavenging.
- Biomedical Pioneering Innovation Center (BIOPIC), and State Key Laboratory of Gene Function and Modulation Research, School of Life Sciences, Peking University, Beijing, China. xuhua@pku.edu.cn.
Organizational Affiliation: