Structure of the TOM40 complex annealed
Yang, L.Y., Shen, Q.T.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mitochondrial import receptor subunit TOM22 | A [auth B], F [auth G] | 172 | Saccharomyces cerevisiae | Mutation(s): 0 Gene Names: TOM22, MAS17, MAS22, MOM22, YNL131W, N1217, N1862 Membrane Entity: Yes |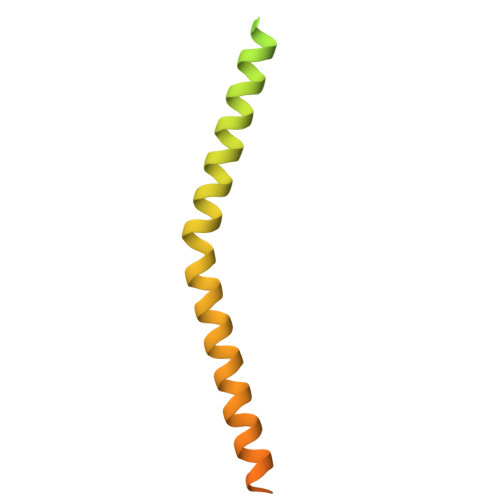  |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P49334 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mitochondrial import receptor subunit TOM5 | B [auth C], G [auth H] | 50 | Saccharomyces cerevisiae | Mutation(s): 0 Membrane Entity: Yes |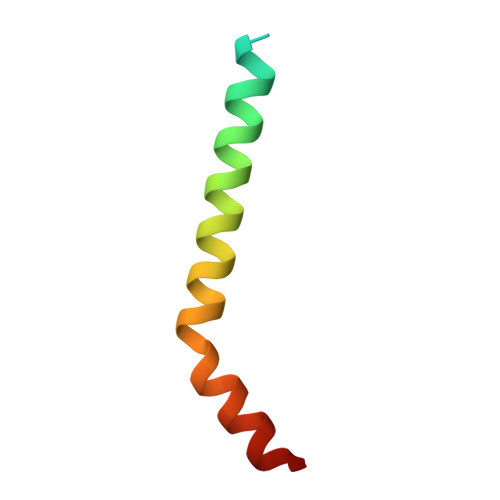  |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P80967 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mitochondrial import receptor subunit TOM6 | C [auth D], H [auth I] | 61 | Saccharomyces cerevisiae | Mutation(s): 0 Membrane Entity: Yes |  |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P33448 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mitochondrial import receptor subunit TOM7 | D [auth E], I [auth J] | 60 | Saccharomyces cerevisiae | Mutation(s): 0 Membrane Entity: Yes |  |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P53507 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mitochondrial import receptor subunit TOM40 | E [auth A], J [auth F] | 387 | Saccharomyces cerevisiae | Mutation(s): 0 Membrane Entity: Yes |  |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P23644 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Science Foundation (NSF, China) | China | -- |