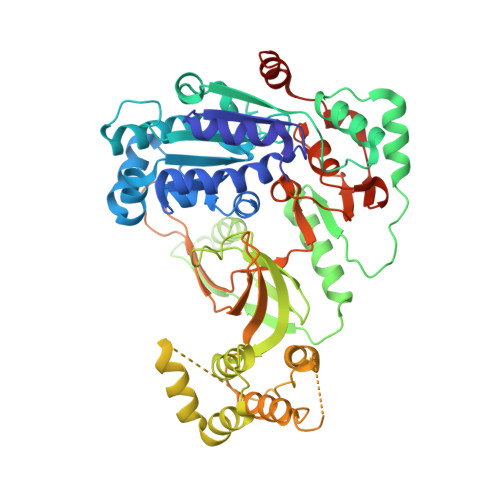
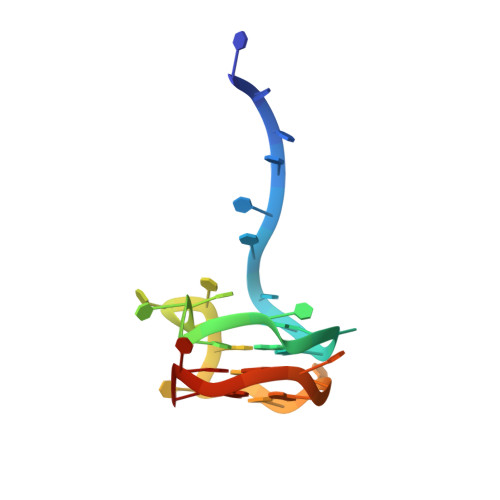
Eukaryotic Pif1 helicase unwinds G-quadruplex and dsDNA using a conserved wedge.
Hong, Z., Byrd, A.K., Gao, J., Das, P., Tan, V.Q., Malone, E.G., Osei, B., Marecki, J.C., Protacio, R.U., Wahls, W.P., Raney, K.D., Song, H.(2024) Nat Commun 15: 6104-6104
- PubMed: 39030241 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-50575-8
- Primary Citation Related Structures:
8XAK - PubMed Abstract:
G-quadruplexes (G4s) formed by guanine-rich nucleic acids induce genome instability through impeding DNA replication fork progression. G4s are stable DNA structures, the unfolding of which require the functions of DNA helicases. Pif1 helicase binds preferentially to G4 DNA and plays multiple roles in maintaining genome stability, but the mechanism by which Pif1 unfolds G4s is poorly understood. Here we report the co-crystal structure of Saccharomyces cerevisiae Pif1 (ScPif1) bound to a G4 DNA with a 5' single-stranded DNA (ssDNA) segment. Unlike the Thermus oshimai Pif1-G4 structure, in which the 1B and 2B domains confer G4 recognition, ScPif1 recognizes G4 mainly through the wedge region in the 1A domain that contacts the 5' most G-tetrad directly. A conserved Arg residue in the wedge is required for Okazaki fragment processing but not for mitochondrial function or for suppression of gross chromosomal rearrangements. Multiple substitutions at this position have similar effects on resolution of DNA duplexes and G4s, suggesting that ScPif1 may use the same wedge to unwind G4 and dsDNA. Our results reveal the mechanism governing dsDNA unwinding and G4 unfolding by ScPif1 helicase that can potentially be generalized to other eukaryotic Pif1 helicases and beyond.
- Institute of Molecular and Cell Biology (IMCB), Agency for Science, Technology and Research (A*STAR), Proteos, Singapore, Republic of Singapore.
Organizational Affiliation: