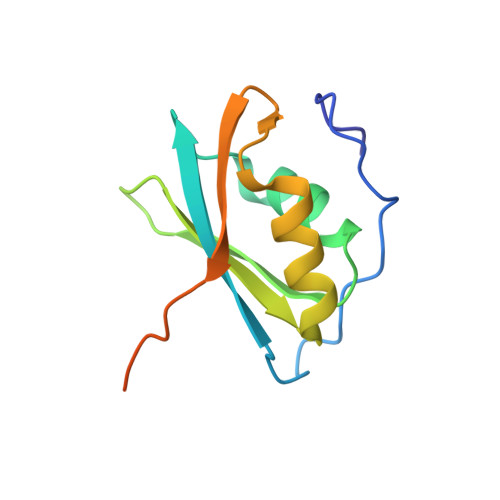
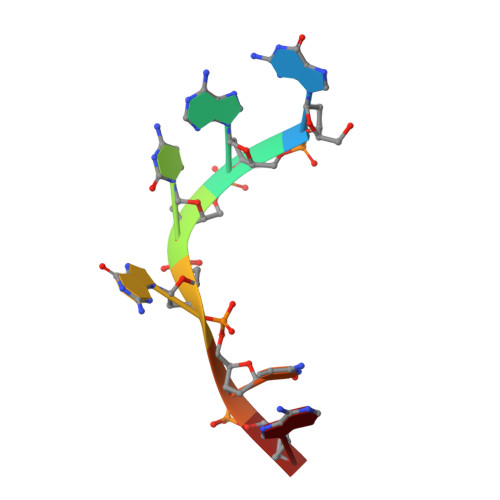
Structural basis for RNA recognition by the C-terminal RRM domain of human RBM45.
Chen, X., Wei, Q., Yang, Z., Chen, X., Guo, S., Jiang, M., Wang, M.(2024) J Biological Chem 300: 107640-107640
- PubMed: 39122006 Search on PubMed
- DOI: https://doi.org/10.1016/j.jbc.2024.107640
- Primary Citation Related Structures:
8WQ3, 8WQ5 - PubMed Abstract:
RBM45 is an RNA-binding protein with roles in neural development by regulating RNA splicing. Its dysfunction and aggregation are associated with neurodegenerative diseases such as amyotrophic lateral sclerosis (ALS) and frontotemporal lobar dementia (FTLD). RBM45 harbors three RRM domains that potentially bind RNA. While the recognitions of RNA by its N-terminal tandem RRM domains (RRM1 and RRM2) have been well understood, the RNA-binding property of its C-terminal RRM (RRM3) remains unclear. In this work, we identified that the RRM3 of RBM45 sequence-specifically binds RNA with a GACG sequence, similar but not identical to those recognized by the RRM1 and RRM2. Further, we determined the crystal structure of RBM45 RRM3 in complex with a GACG sequence-containing single-stranded DNA. Our structural results, together with the RNA-binding assays of mutants at key amino acid residues, revealed the molecular mechanism by which RBM45 RRM3 recognizes an RNA sequence. Our finding on the RNA-binding property of the individual RRM module of RBM45 provides the foundation for unraveling the RNA-binding characteristics of full-length RBM45 and for understanding the biological functions of RBM45.
- Institutes of Physical Science and Information Technology, Anhui University, Hefei 230601, Anhui, China; School of Life Sciences, Anhui University, Hefei 230601, Anhui, China.
Organizational Affiliation: