Structure of Cul2-VCB-Protac-Wee1 complex at 3.6 Angstrom resolution.
Wang, P., Zhang, T.T.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cullin-2 | 745 | Homo sapiens | Mutation(s): 0 Gene Names: CUL2 | 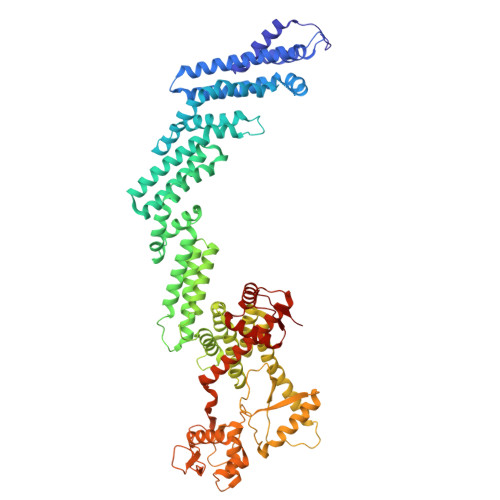 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q13617 GTEx: ENSG00000108094 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q13617 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| E3 ubiquitin-protein ligase RBX1, N-terminally processed | B [auth R] | 82 | Homo sapiens | Mutation(s): 0 Gene Names: RBX1 EC: 2.3.2.32 (UniProt), 2.3.2.27 (UniProt) | 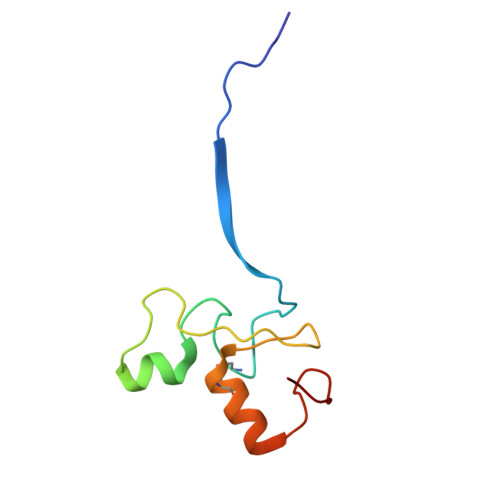 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P62877 GTEx: ENSG00000100387 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P62877 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| von Hippel-Lindau disease tumor suppressor | C [auth V] | 155 | Homo sapiens | Mutation(s): 0 Gene Names: VHL |  |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P40337 GTEx: ENSG00000134086 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P40337 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Elongin-B | D [auth B] | 104 | Homo sapiens | Mutation(s): 0 Gene Names: ELOB, TCEB2 | 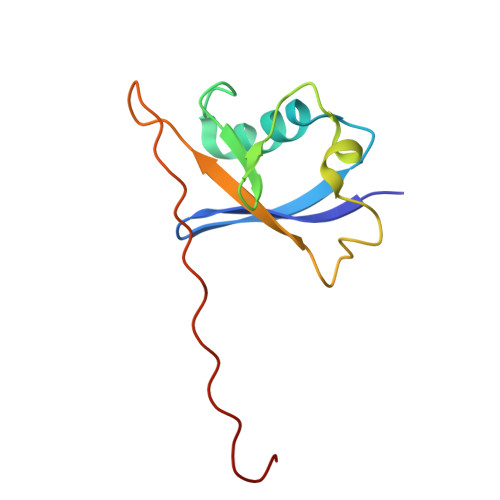 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q15370 GTEx: ENSG00000103363 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q15370 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Elongin-C | E [auth C] | 96 | Homo sapiens | Mutation(s): 0 Gene Names: ELOC | 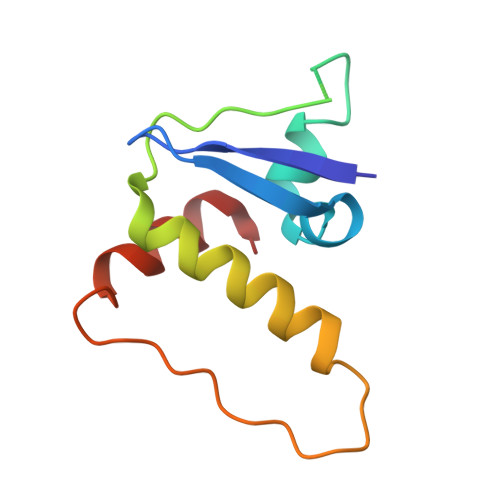 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q15369 GTEx: ENSG00000154582 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q15369 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 6 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Wee1-like protein kinase | F [auth W] | 286 | Homo sapiens | Mutation(s): 0 Gene Names: WEE1 EC: 2.7.10.2 |  |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P30291 GTEx: ENSG00000166483 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P30291 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| W6U (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | G [auth V] | (2S,4R)-1-[(2S)-3,3-dimethyl-2-[3-[4-[4-[4-[[3-oxidanylidene-1-[6-(2-oxidanylpropan-2-yl)pyridin-2-yl]-2-prop-2-enyl-pyrazolo[3,4-d]pyrimidin-6-yl]amino]phenyl]piperazin-1-yl]butoxy]propanoylamino]butanoyl]-N-[[4-(4-methyl-1,3-thiazol-5-yl)phenyl]methyl]-4-oxidanyl-pyrrolidine-2-carboxamide C55 H70 N12 O7 S QQLAKZIDPLGLSV-VZUPZQALSA-N |  | ||
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |