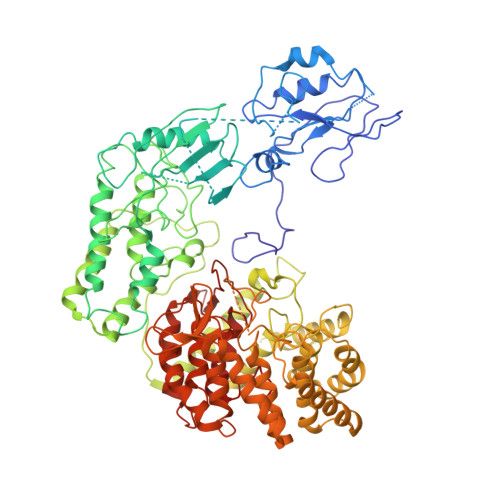
Mechanistic insights into lanthipeptide modification by a distinct subclass of LanKC enzyme that forms dimers.
Li, Y., Shao, K., Li, Z., Zhu, K., Gan, B.K., Shi, J., Xiao, Y., Luo, M.(2024) Nat Commun 15: 7090-7090
- PubMed: 39154050 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-51600-6
- Primary Citation Related Structures:
8W7A, 8W7J, 8WGO - PubMed Abstract:
Naturally occurring lanthipeptides, peptides post-translationally modified by various enzymes, hold significant promise as antibiotics. Despite extensive biochemical and structural studies, the events preceding peptide modification remain poorly understood. Here, we identify a distinct subclass of lanthionine synthetase KC (LanKC) enzymes with distinct structural and functional characteristics. We show that PneKC, a member of this subclass, forms a dimer and possesses GTPase activity. Through three cryo-EM structures of PneKC, we illustrate different stages of peptide PneA binding, from initial recognition to full binding. Our structures show the kinase domain complexed with the PneA core peptide and GTPγS, a phosphate-bound lyase domain, and an unconventional cyclase domain. The leader peptide of PneA interact with a gate loop, transitioning from an extended to a helical conformation. We identify a dimerization hot spot and propose a "negative cooperativity" mechanism toggling the enzyme between tense and relaxed conformation. Additionally, we identify an important salt bridge in the cyclase domain, differing from those in in conventional cyclase domains. These residues are highly conserved in the LanKC subclass and are part of two signature motifs. These results unveil potential differences in lanthipeptide modification enzymes assembly and deepen our understanding of allostery in these multifunctional enzymes.
- Department of Biological sciences, Faculty of Science, National University of Singapore, Singapore, Singapore.
Organizational Affiliation: