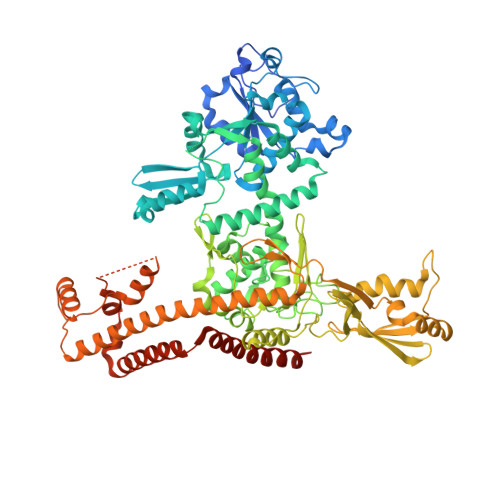
Structural insights into the assembly of type IIA topoisomerase DNA cleavage-religation center.
Liu, K.T., Chen, S.F., Chan, N.L.(2024) Nucleic Acids Res 52: 9788-9802
- PubMed: 39077950 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkae657
- Primary Citation Related Structures:
8KE7, 8W50 - PubMed Abstract:
The ability to catalyze reversible DNA cleavage and religation is central to topoisomerases' role in regulating DNA topology. In type IIA topoisomerases (Top2), the formation of its DNA cleavage-religation center is driven by DNA-binding-induced structural rearrangements. These changes optimally position key catalytic modules, such as the active site tyrosine of the WHD domain and metal ion(s) chelated by the TOPRIM domain, around the scissile phosphodiester bond to perform reversible transesterification. To understand this assembly process in detail, we report the catalytic core structures of human Top2α and Top2β in an on-pathway conformational state. This state features an in trans formation of an interface between the Tower and opposing TOPRIM domain, revealing a groove for accommodating incoming G-segment DNA. Structural superimposition further unveils how subsequent DNA-binding-induced disengagement of the TOPRIM and Tower domains allows a firm grasp of the bound DNA for cleavage/religation. Notably, we identified a previously undocumented protein-DNA interaction, formed between an arginine-capped C-terminus of an α-helix in the TOPRIM domain and the DNA backbone, significantly contributing to Top2 function. This work uncovers a previously unrecognized role of the Tower domain, highlighting its involvement in anchoring and releasing the TOPRIM domain, thus priming Top2 for DNA binding and cleavage.
- Institute of Biochemistry and Molecular Biology, College of Medicine, National Taiwan University, Taipei 100, Taiwan.
Organizational Affiliation: