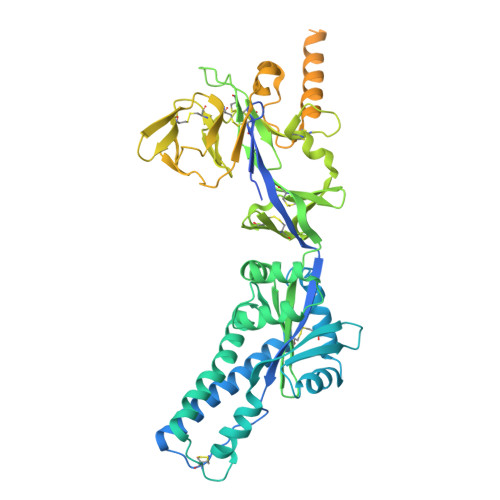
Rational design of uncleaved prefusion-closed trimer vaccines for human respiratory syncytial virus and metapneumovirus.
Lee, Y.Z., Han, J., Zhang, Y.N., Ward, G., Braz Gomes, K., Auclair, S., Stanfield, R.L., He, L., Wilson, I.A., Zhu, J.(2024) Nat Commun 15: 9939-9939
- PubMed: 39550381 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-54287-x
- Primary Citation Related Structures:
8W3E, 8W3F, 8W3G, 8W3H, 8W3I, 8W3J, 8W3K, 8W3L, 8W3N, 8W3O, 8W3P, 8W3Q, 8W3R - PubMed Abstract:
Respiratory syncytial virus (RSV) and human metapneumovirus (hMPV) cause human respiratory diseases and are major targets for vaccine development. In this study, we design uncleaved prefusion-closed (UFC) trimers for the fusion protein (F) of both viruses by examining mutations critical to F metastability. For RSV, we assess four previous prefusion F designs, including the first and second generations of DS-Cav1, SC-TM, and 847A. We then identify key mutations that can maintain prefusion F in a native-like, closed trimeric form (up to 76%) without introducing any interprotomer disulfide bond. For hMPV, we develop a stable UFC trimer with a truncated F 2 -F 1 linkage and an interprotomer disulfide bond. Dozens of UFC constructs are characterized by negative-stain electron microscopy (nsEM), x-ray crystallography (11 RSV-F structures and one hMPV-F structure), and antigenic profiling. Using an optimized RSV-F UFC trimer as bait, we identify three potent RSV neutralizing antibodies (NAbs) from a phage-displayed human antibody library, with a public NAb lineage targeting sites Ø and V and two cross-pneumovirus NAbs recognizing site III. In mouse immunization, rationally designed RSV-F and hMPV-F UFC trimers induce robust antibody responses with high neutralizing titers. Our study provides a foundation for future prefusion F-based RSV and hMPV vaccine development.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA, 92037, USA.
Organizational Affiliation: