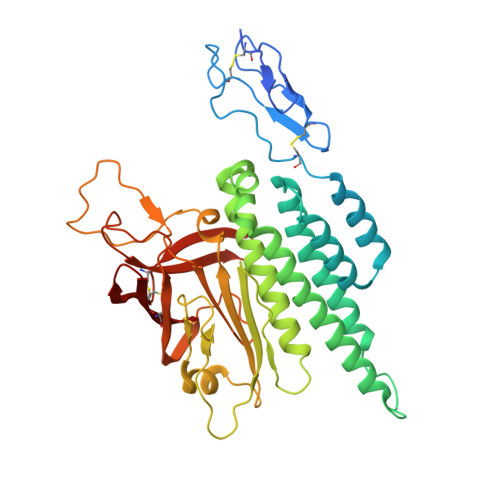
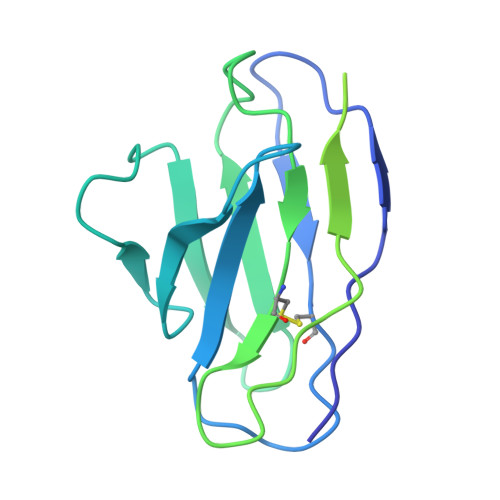
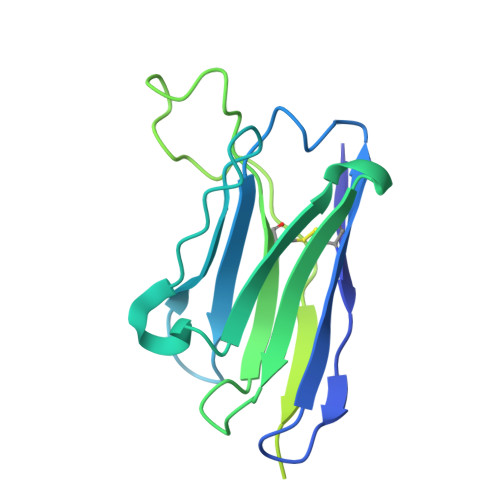
Conformational coupling between extracellular and transmembrane domains modulates holo-adhesion GPCR function.
Kordon, S.P., Cechova, K., Bandekar, S.J., Leon, K., Dutka, P., Siffer, G., Kossiakoff, A.A., Vafabakhsh, R., Arac, D.(2024) Nat Commun 15: 10545-10545
- PubMed: 39627215 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-54836-4
- Primary Citation Related Structures:
8VTI - PubMed Abstract:
Adhesion G Protein-Coupled Receptors (aGPCRs) are key cell-adhesion molecules involved in numerous physiological functions. aGPCRs have large multi-domain extracellular regions (ECRs) containing a conserved GAIN domain that precedes their seven-pass transmembrane domain (7TM). Ligand binding and mechanical force applied on the ECR regulate receptor function. However, how the ECR communicates with the 7TM remains elusive, because the relative orientation and dynamics of the ECR and 7TM within a holoreceptor is unclear. Here, we describe the cryo-EM reconstruction of an aGPCR, Latrophilin3/ADGRL3, and reveal that the GAIN domain adopts a parallel orientation to the transmembrane region and has constrained movement. Single-molecule FRET experiments unveil three slow-exchanging FRET states of the ECR relative to the transmembrane region within the holoreceptor. GAIN-targeted antibodies, and cancer-associated mutations at the GAIN-7TM interface, alter FRET states, cryo-EM conformations, and receptor signaling. Altogether, this data demonstrates conformational and functional coupling between the ECR and 7TM, suggesting an ECR-mediated mechanism for aGPCR activation.
- Department of Biochemistry and Molecular Biology, The University of Chicago, Chicago, IL, USA.
Organizational Affiliation: