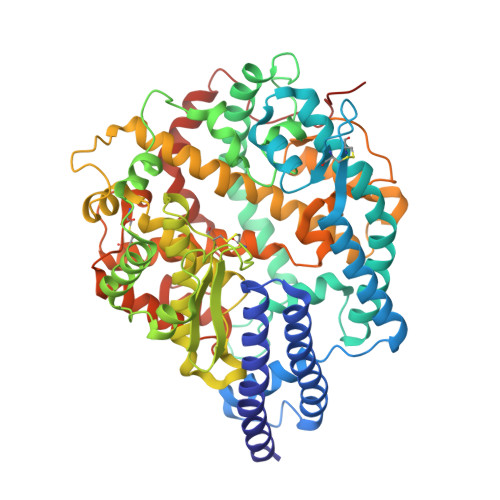
Structural basis for raccoon dog receptor recognition by SARS-CoV-2.
Hsueh, F.C., Shi, K., Mendoza, A., Bu, F., Zhang, W., Aihara, H., Li, F.(2024) PLoS Pathog 20: e1012204-e1012204
- PubMed: 38709834 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1012204
- Primary Citation Related Structures:
8VQR - PubMed Abstract:
Since the COVID-19 outbreak, raccoon dogs have been suggested as a potential intermediary in transmitting SARS-CoV-2 to humans. To understand their role in the COVID-19 pandemic and the species barrier for SARS-CoV-2 transmission to humans, we analyzed how their ACE2 protein interacts with SARS-CoV-2 spike protein. Biochemical data showed that raccoon dog ACE2 is an effective receptor for SARS-CoV-2 spike protein, though not as effective as human ACE2. Structural comparisons highlighted differences in the virus-binding residues of raccoon dog ACE2 compared to human ACE2 (L24Q, Y34H, E38D, T82M, R353K), explaining their varied effectiveness as receptors for SARS-CoV-2. These variations contribute to the species barrier that exists between raccoon dogs and humans regarding SARS-CoV-2 transmission. Identifying these barriers can help assess the susceptibility of other mammals to SARS-CoV-2. Our research underscores the potential of raccoon dogs as SARS-CoV-2 carriers and identifies molecular barriers that affect the virus's ability to jump between species.
- Department of Pharmacology, University of Minnesota Medical School, Minneapolis, Minnesota, United States of America.
Organizational Affiliation: