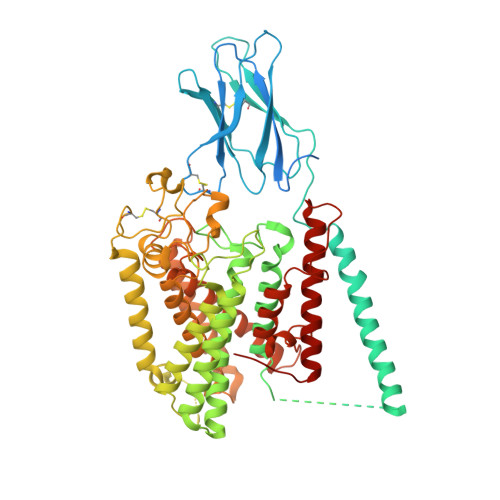
Structural and mechanistic insights into a lysosomal membrane enzyme HGSNAT involved in Sanfilippo syndrome.
Zhao, B., Cao, Z., Zheng, Y., Nguyen, P., Bowen, A., Edwards, R.H., Stroud, R.M., Zhou, Y., Van Lookeren Campagne, M., Li, F.(2024) Nat Commun 15: 5388-5388
- PubMed: 38918376 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-49614-1
- Primary Citation Related Structures:
8VKJ, 8VLG, 8VLI, 8VLU, 8VLV, 8VLY - PubMed Abstract:
Heparan sulfate (HS) is degraded in lysosome by a series of glycosidases. Before the glycosidases can act, the terminal glucosamine of HS must be acetylated by the integral lysosomal membrane enzyme heparan-α-glucosaminide N-acetyltransferase (HGSNAT). Mutations of HGSNAT cause HS accumulation and consequently mucopolysaccharidosis IIIC, a devastating lysosomal storage disease characterized by progressive neurological deterioration and early death where no treatment is available. HGSNAT catalyzes a unique transmembrane acetylation reaction where the acetyl group of cytosolic acetyl-CoA is transported across the lysosomal membrane and attached to HS in one reaction. However, the reaction mechanism remains elusive. Here we report six cryo-EM structures of HGSNAT along the reaction pathway. These structures reveal a dimer arrangement and a unique structural fold, which enables the elucidation of the reaction mechanism. We find that a central pore within each monomer traverses the membrane and controls access of cytosolic acetyl-CoA to the active site at its luminal mouth where glucosamine binds. A histidine-aspartic acid catalytic dyad catalyzes the transfer reaction via a ternary complex mechanism. Furthermore, the structures allow the mapping of disease-causing variants and reveal their potential impact on the function, thus creating a framework to guide structure-based drug discovery efforts.
- Amgen Research, Department of Structural biology, South San Francisco, CA, USA.
Organizational Affiliation: