Co-crystal structure of human TREX1 in complex with an inhibitor
Dehghani-Tafti, S., Dong, A., Li, Y., Ackloo, S., Arrowsmith, C.H., Edwards, A.M., Halabelian, L.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Three-prime repair exonuclease 1 | 233 | Homo sapiens | Mutation(s): 0 Gene Names: TREX1 EC: 3.1.11.2 | 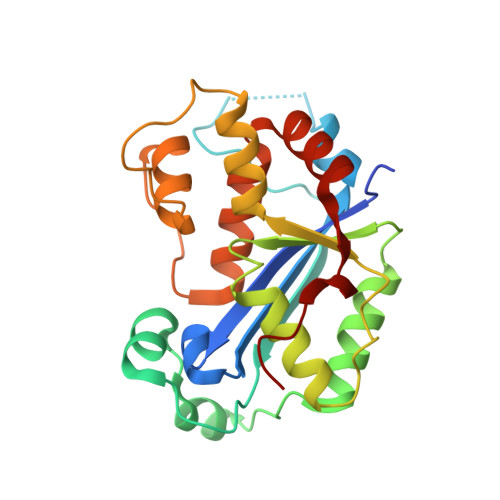 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9NSU2 GTEx: ENSG00000213689 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9NSU2 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| A1ACJ (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | AA [auth E] EA [auth F] I [auth A] IA [auth G] MA [auth H] | (2P)-2-[3-bromo-2-(2-hydroxyethoxy)phenyl]-5-hydroxy-1-methyl-N-(1,2-oxazol-4-yl)-6-oxo-1,6-dihydropyrimidine-4-carboxamide C17 H15 Br N4 O6 HNZOETDJOULZPF-UHFFFAOYSA-N |  | ||
| PG4 Download:Ideal Coordinates CCD File | DA [auth E], V [auth C] | TETRAETHYLENE GLYCOL C8 H18 O5 UWHCKJMYHZGTIT-UHFFFAOYSA-N |  | ||
| PEG Download:Ideal Coordinates CCD File | W [auth C] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | J [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| MG Download:Ideal Coordinates CCD File | BA [auth E] CA [auth E] FA [auth F] GA [auth F] JA [auth G] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| UNX Download:Ideal Coordinates CCD File | HA [auth F], LA [auth G], M [auth A], N [auth A], R [auth B] | UNKNOWN ATOM OR ION X |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 55.827 | α = 93.12 |
| b = 56.336 | β = 98.22 |
| c = 162.853 | γ = 99.92 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| HKL-3000 | data scaling |
| HKL-3000 | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Other private | -- |