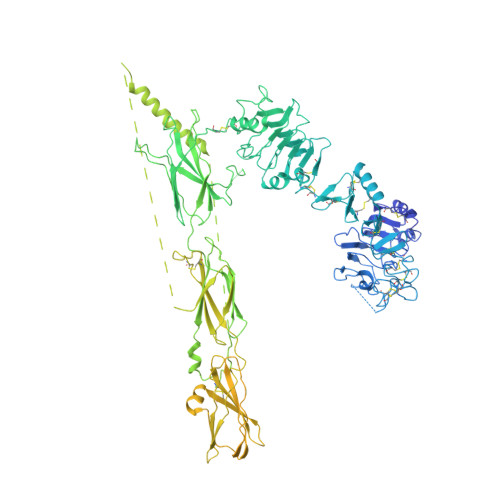
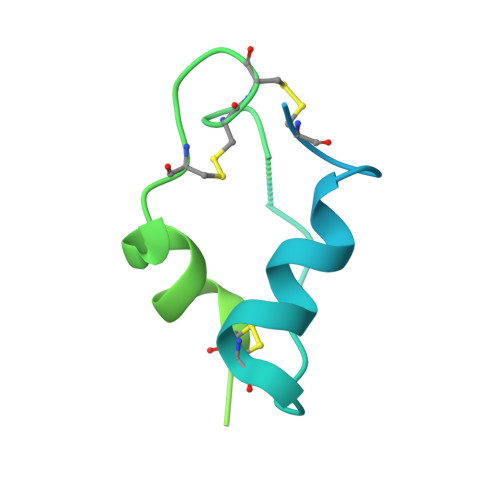
Activation of the insulin receptor by insulin-like growth factor 2.
An, W., Hall, C., Li, J., Hung, A., Wu, J., Park, J., Wang, L., Bai, X.C., Choi, E.(2024) Nat Commun 15: 2609-2609
- PubMed: 38521788 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-46990-6
- Primary Citation Related Structures:
8U4B, 8U4C, 8U4E, 8VJB, 8VJC - PubMed Abstract:
Insulin receptor (IR) controls growth and metabolism. Insulin-like growth factor 2 (IGF2) has different binding properties on two IR isoforms, mimicking insulin's function. However, the molecular mechanism underlying IGF2-induced IR activation remains unclear. Here, we present cryo-EM structures of full-length human long isoform IR (IR-B) in both the inactive and IGF2-bound active states, and short isoform IR (IR-A) in the IGF2-bound active state. Under saturated IGF2 concentrations, both the IR-A and IR-B adopt predominantly asymmetric conformations with two or three IGF2s bound at site-1 and site-2, which differs from that insulin saturated IR forms an exclusively T-shaped symmetric conformation. IGF2 exhibits a relatively weak binding to IR site-2 compared to insulin, making it less potent in promoting full IR activation. Cell-based experiments validated the functional importance of IGF2 binding to two distinct binding sites in optimal IR signaling and trafficking. In the inactive state, the C-terminus of α-CT of IR-B contacts FnIII-2 domain of the same protomer, hindering its threading into the C-loop of IGF2, thus reducing the association rate of IGF2 with IR-B. Collectively, our studies demonstrate the activation mechanism of IR by IGF2 and reveal the molecular basis underlying the different affinity of IGF2 to IR-A and IR-B.
- Department of Biophysics, University of Texas Southwestern Medical Center, Dallas, TX, 75390, USA.
Organizational Affiliation: