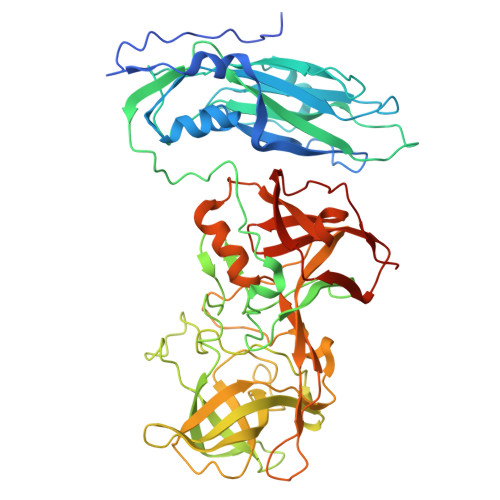
Antibody-Based Affinity Cryoelectron Microscopy at 2.6- angstrom Resolution.
Yu, G., Li, K., Huang, P., Jiang, X., Jiang, W.(2016) Structure 24: 1984-1990
- PubMed: 27806259 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2016.09.008
- Primary Citation Related Structures:
8VG6 - PubMed Abstract:
The affinity cryoelectron microscopy (cryo-EM) approach has been explored in recent years to simplify and/or improve the sample preparation for cryo-EM, which can bring previously challenging specimens such as those of low abundance and/or unpurified ones within reach of the cryo-EM technique. Despite the demonstrated successes for solving structures to low to intermediate resolutions, the lack of near-atomic structures using this approach has led to a common perception of affinity cryo-EM as a niche technique incapable of reaching high resolutions. Here, we report a ∼2.6-Å structure solved using the antibody-based affinity grid approach with low-concentration Tulane virus purified from a low-yield cell-culture system that has been challenging to standard cryo-EM grid preparation. Quantitative analyses of the structure indicate data and reconstruction quality comparable with the conventional grid preparation method using samples at high concentration.
- Department of Biological Science, Markey Center for Structural Biology, Purdue University, West Lafayette, IN 47907, USA.
Organizational Affiliation: