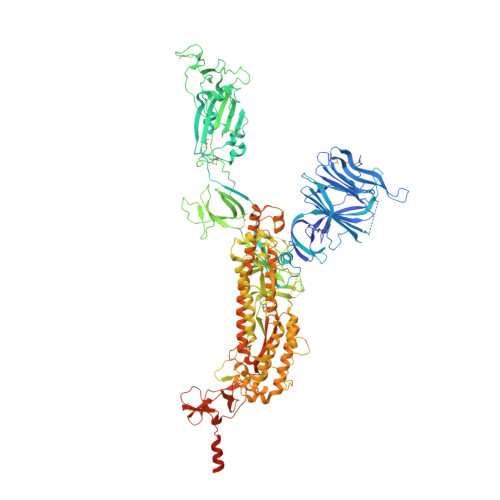
Lys417 acts as a molecular switch that regulates the conformation of SARS-CoV-2 spike protein.
Geng, Q., Wan, Y., Hsueh, F.C., Shang, J., Ye, G., Bu, F., Herbst, M., Wilkens, R., Liu, B., Li, F.(2023) Elife 12
- PubMed: 37991488 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.74060
- Primary Citation Related Structures:
8UUL, 8UUM, 8UUN, 8UUO - PubMed Abstract:
SARS-CoV-2 spike protein plays a key role in mediating viral entry and inducing host immune responses. It can adopt either an open or closed conformation based on the position of its receptor-binding domain (RBD). It is yet unclear what causes these conformational changes or how they influence the spike's functions. Here, we show that Lys417 in the RBD plays dual roles in the spike's structure: it stabilizes the closed conformation of the trimeric spike by mediating inter-spike-subunit interactions; it also directly interacts with ACE2 receptor. Hence, a K417V mutation has opposing effects on the spike's function: it opens up the spike for better ACE2 binding while weakening the RBD's direct binding to ACE2. The net outcomes of this mutation are to allow the spike to bind ACE2 with higher probability and mediate viral entry more efficiently, but become more exposed to neutralizing antibodies. Given that residue 417 has been a viral mutational hotspot, SARS-CoV-2 may have been evolving to strike a balance between infection potency and immune evasion, contributing to its pandemic spread.
- Department of Pharmacology, University of Minnesota Medical School, Minneapolis, United States.
Organizational Affiliation: