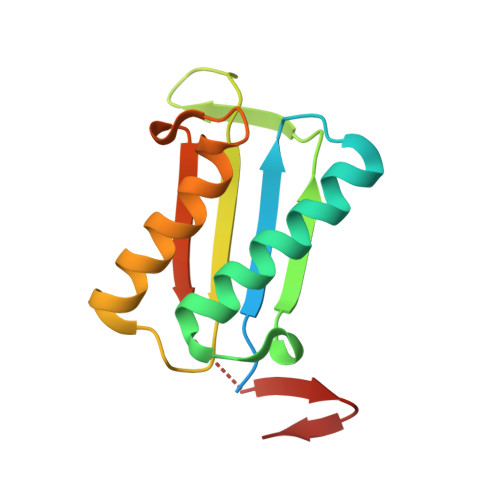
Structures of Trichomonas vaginalis macrophage migratory inhibitory factor.
Srivastava, A., Nair, A., Dawson, O.C.O., Gao, R., Liu, L., Craig, J.K., Battaile, K.P., Harmon, E.K., Barrett, L.K., Van Voorhis, W.C., Subramanian, S., Myler, P.J., Lovell, S., Asojo, O.A., Darwiche, R.(2024) Acta Crystallogr F Struct Biol Commun 80: 341-347
- PubMed: 39601418 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X24011105
- Primary Citation Related Structures:
8UR2, 8UR4, 8UZ4 - PubMed Abstract:
The unicellular parasitic protozoan Trichomonas vaginalis causes trichomoniasis, the most prevalent nonviral sexually transmitted disease globally. T. vaginalis evades host immune responses by producing homologs of host proteins, including cytokines such as macrophage migration inhibitory factor. T. vaginalis macrophage migration inhibitory factor (TvMIF) helps to facilitate the survival of T. vaginalis during nutritional stress conditions, increases prostate cell proliferation and invasiveness, and induces inflammation-related cellular pathways, thus mimicking the ability of human MIF to increase inflammation and cell proliferation. The production, crystallization and three structures of N-terminally hexahistidine-tagged TvMIF reveal a prototypical MIF trimer with a topology similar to that of human homologs (hMIF-1 and hMIF-2). The N-terminal tag obscures the expected pyruvate-binding site. The similarity of TvMIF to its human homologs can be exploited for structure-based drug discovery.
- California Institute of Technology, 1200 East California Boulevard, Pasadena, CA 91125, USA.
Organizational Affiliation: