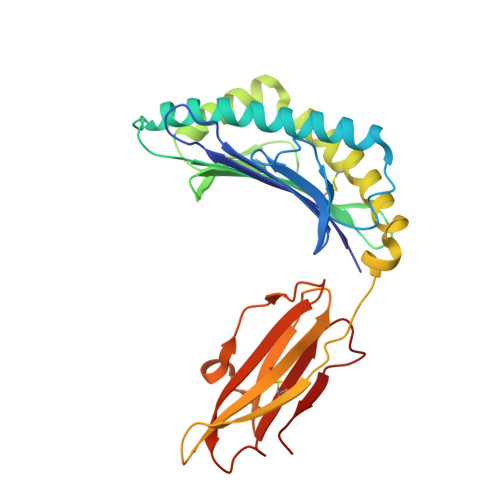
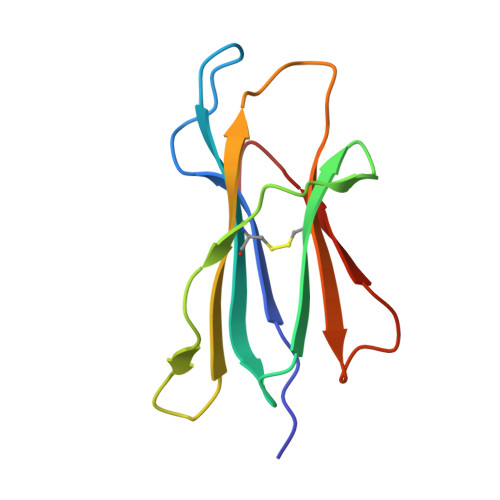
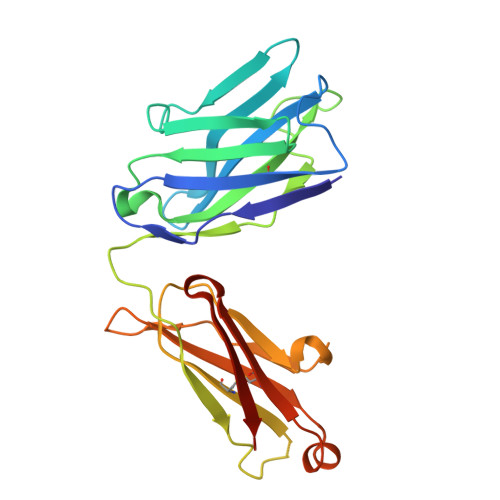
Therapeutic Targeting and Structural Characterization of a Sotorasib-Modified KRAS G12C-MHC I Complex Demonstrate the Antitumor Efficacy of Hapten-Based Strategies.
Pandey, A., Rohweder, P.J., Chan, L.M., Ongpipattanakul, C., Chung, D.H., Paolella, B., Quimby, F.M., Nguyen, N., Verba, K.A., Evans, M.J., Craik, C.S.(2025) Cancer Res 85: 329-341
- PubMed: 39656104 Search on PubMed
- DOI: https://doi.org/10.1158/0008-5472.CAN-24-2450
- Primary Citation Related Structures:
8UDR - PubMed Abstract:
Antibody-based therapies have emerged as a powerful strategy for the management of diverse cancers. Unfortunately, tumor-specific antigens remain challenging to identify and target. Recent work established that inhibitor-modified peptide adducts derived from KRAS G12C are competent for antigen presentation via MHC I and can be targeted by antibody-based therapeutics, offering a means to directly target an intracellular oncoprotein at the cell surface with combination therapies. Here, we validated antigen display of "haptenated" KRAS G12C peptide fragments on tumors in mouse models treated with the FDA-approved KRAS G12C covalent inhibitor Sotorasib using PET/CT imaging of an 89Zr-labeled P1B7 IgG antibody, which selectively binds Sotorasib-modified KRAS G12C MHC I complexes. Targeting this peptide-MHC I complex with radioligand therapy using 225Ac- or 177Lu-P1B7 IgG effectively inhibited tumor growth in combination with Sotorasib. Elucidation of the 3.1 Å cryo-EM structure of P1B7 bound to a haptenated KRAS G12C peptide-MHC I complex confirmed that the Sotorasib-modified KRAS G12C peptide is presented via a canonical binding pose and showed that P1B7 binds the complex in a T-cell receptor-like manner. Together, these findings demonstrate the potential value of targeting unique oncoprotein-derived, haptenated MHC I complexes with radioligand therapeutics and provide a structural framework for developing next generation antibodies.
- University of California, San Francisco, San Francisco, United States.
Organizational Affiliation: