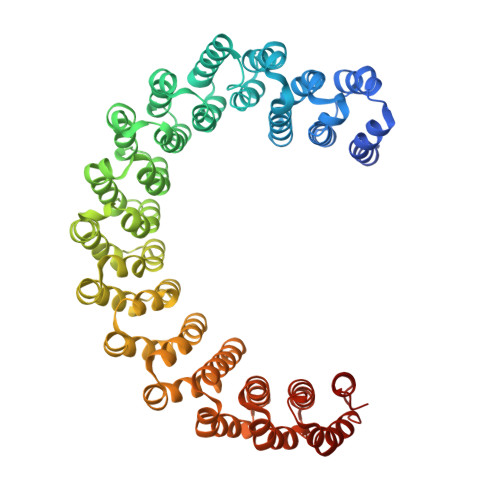B56 delta long-disordered arms form a dynamic PP2A regulation interface coupled with global allostery and Jordan's syndrome mutations.
Wu, C.G., Balakrishnan, V.K., Merrill, R.A., Parihar, P.S., Konovolov, K., Chen, Y.C., Xu, Z., Wei, H., Sundaresan, R., Cui, Q., Wadzinski, B.E., Swingle, M.R., Musiyenko, A., Chung, W.K., Honkanen, R.E., Suzuki, A., Huang, X., Strack, S., Xing, Y.(2024) Proc Natl Acad Sci U S A 121: e2310727120-e2310727120
- PubMed: 38150499 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2310727120
- Primary Citation Related Structures:
8U1X, 8U89 - PubMed Abstract:
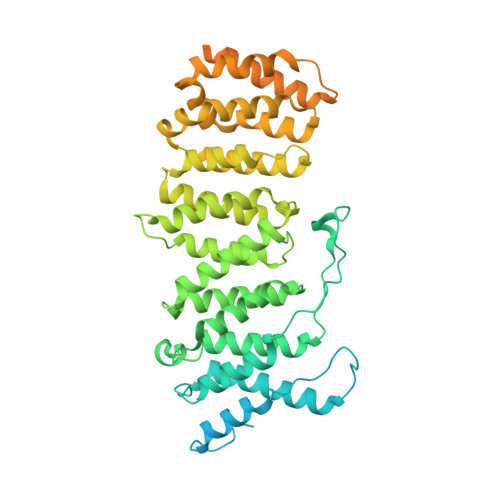
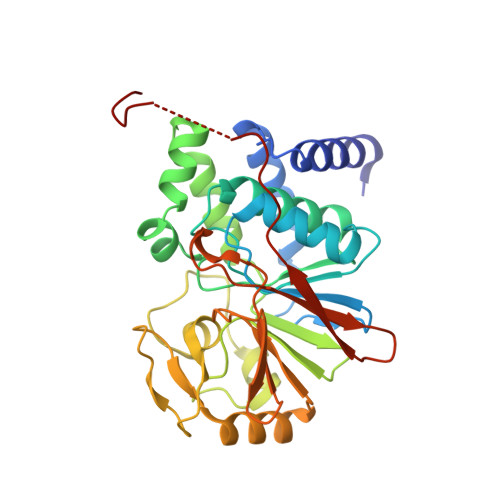
Intrinsically disordered regions (IDR) and short linear motifs (SLiMs) play pivotal roles in the intricate signaling networks governed by phosphatases and kinases. B56δ (encoded by PPP2R5D ) is a regulatory subunit of protein phosphatase 2A (PP2A) with long IDRs that harbor a substrate-mimicking SLiM and multiple phosphorylation sites. De novo missense mutations in PPP2R5D cause intellectual disabilities (ID), macrocephaly, Parkinsonism, and a broad range of neurological symptoms. Our single-particle cryo-EM structures of the PP2A-B56δ holoenzyme reveal that the long, disordered arms at the B56δ termini fold against each other and the holoenzyme core. This architecture suppresses both the phosphatase active site and the substrate-binding protein groove, thereby stabilizing the enzyme in a closed latent form with dual autoinhibition. The resulting interface spans over 190 Å and harbors unfavorable contacts, activation phosphorylation sites, and nearly all residues with ID-associated mutations. Our studies suggest that this dynamic interface is coupled to an allosteric network responsive to phosphorylation and altered globally by mutations. Furthermore, we found that ID mutations increase the holoenzyme activity and perturb the phosphorylation rates, and the severe variants significantly increase the mitotic duration and error rates compared to the normal variant.
- McArdle Laboratory for Cancer Research, Department of Oncology, University of Wisconsin at Madison, School of Medicine and Public Health, Madison, WI 53705.
Organizational Affiliation: