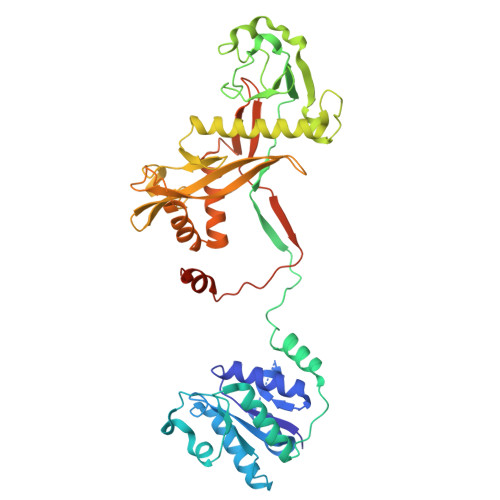
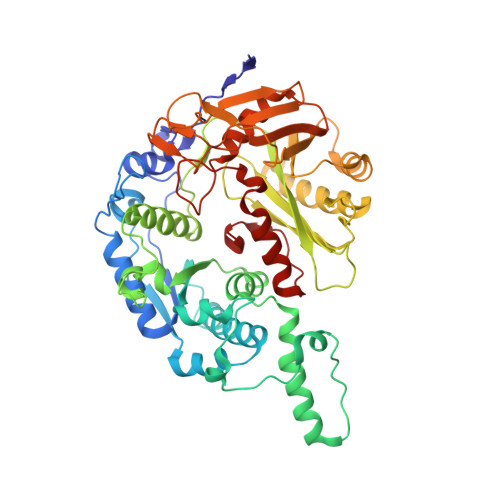
Nucleic acid mediated activation of a short prokaryotic Argonaute immune system.
Kottur, J., Malik, R., Aggarwal, A.K.(2024) Nat Commun 15: 4852-4852
- PubMed: 38844755 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-49271-4
- Primary Citation Related Structures:
8U72, 8U7B - PubMed Abstract:
A short prokaryotic Argonaute (pAgo) TIR-APAZ (SPARTA) defense system, activated by invading DNA to unleash its TIR domain for NAD(P) + hydrolysis, was recently identified in bacteria. We report the crystal structure of SPARTA heterodimer in the absence of guide-RNA/target-ssDNA (2.66 Å) and a cryo-EM structure of the SPARTA oligomer (tetramer of heterodimers) bound to guide-RNA/target-ssDNA at nominal 3.15-3.35 Å resolution. The crystal structure provides a high-resolution view of SPARTA, revealing the APAZ domain as equivalent to the N, L1, and L2 regions of long pAgos and the MID domain containing a unique insertion (insert57). Cryo-EM structure reveals regions of the PIWI (loop10-9) and APAZ (helix αN) domains that reconfigure for nucleic-acid binding and decrypts regions/residues that reorganize to expose a positively charged pocket for higher-order assembly. The TIR domains amass in a parallel-strands arrangement for catalysis. We visualize SPARTA before and after RNA/ssDNA binding and uncover the basis of its active assembly leading to abortive infection.
- Department of Pharmacological Sciences, Icahn School of Medicine at Mount Sinai, New York, NY, 10029, USA. jitheshkottur@iav.res.in.
Organizational Affiliation: