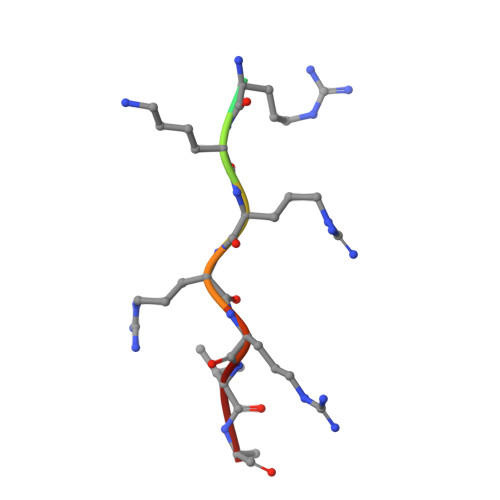
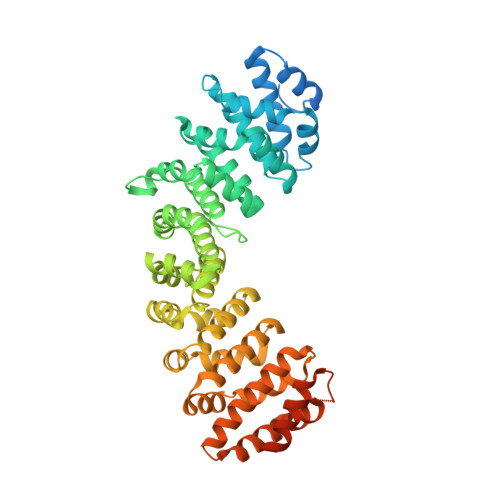Mechanistic Insights Into an Ancient Adenovirus Precursor Protein VII Show Multiple Nuclear Import Receptor Pathways.
Nematollahzadeh, S., Athukorala, A., Donnelly, C.M., Pavan, S., Atelie-Djossou, V., Di Iorio, E., Nath, B., Helbig, K.J., McSharry, B.P., Forwood, J.K., Sarker, S., Alvisi, G.(2024) Traffic 25: e12953-e12953
- PubMed: 39301720 Search on PubMed
- DOI: https://doi.org/10.1111/tra.12953
- Primary Citation Related Structures:
8U36 - PubMed Abstract:
Adenoviral pVII proteins are multifunctional, highly basic, histone-like proteins that can bind to and transport the viral genome into the host cell nucleus. Despite the identification of several nuclear localization signals (NLSs) in the pVII protein of human adenovirus (HAdV)2, the mechanistic details of nuclear transport are largely unknown. Here we provide a full characterization of the nuclear import of precursor (Pre-) pVII protein from an ancient siadenovirus, frog siadenovirus 1 (FrAdV1), using a combination of structural, functional, and biochemical approaches. Two strong NLSs (termed NLSa and NLSd) interact with importin (IMP)β1 and IMPα, respectively, and are the main drivers of nuclear import. A weaker NLS (termed NLSb) also contributes, together with an additional signal (NLSc) which we found to be important for nucleolar targeting and intranuclear binding. Expression of wild-type and NLS defective derivatives Pre-pVII in the presence of selective inhibitors of different nuclear import pathways revealed that, unlike its human counterpart, FrAdV1 Pre-pVII nuclear import is dependent on IMPα/β1 and IMPβ1, but not on transportin-1 (IMPβ2). Clearly, AdVs evolved to maximize the nuclear import pathways for the pVII proteins, whose subcellular localization is the result of a complex process. Therefore, our results pave the way for an evolutionary comparison of the interaction of different AdVs with the host cell nuclear transport machinery.
- Department of Molecular Medicine, University of Padua, Padua, Italy.
Organizational Affiliation: