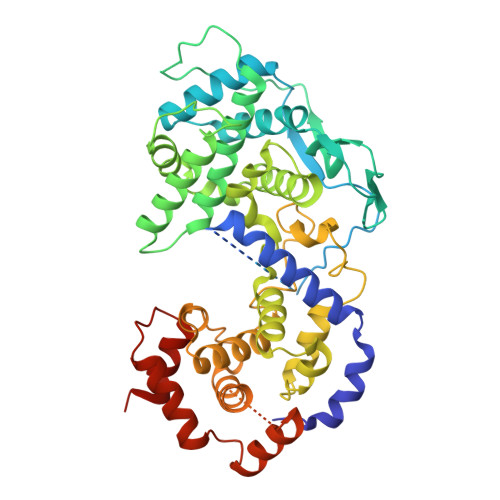
The Replication Function of Rabies Virus P Protein Is Regulated by a Novel Phosphorylation Site in the N-Terminal N Protein-Binding Region
Tudhope, E., Donnelly, C.M., Sethi, A., David, C., Williamson, N., Stewart, M., Forwood, J.K., Gooley, P.R., Moseley, G.W.(2025) Viruses 17