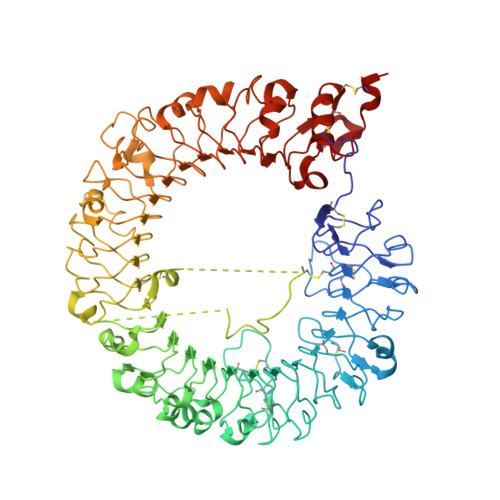
Discovery of Novel TLR7 Agonists as Systemic Agent for Combination With aPD1 for Use in Immuno-oncology.
Poudel, Y.B., He, L., Cox, M., Zhang, Q., Johnson, W.L., Cong, Q., Cheng, H., Chowdari, N.S., Tarby, C., Donnell, A.F., Broekema, M., O'Malley, D.P., Zhang, Y., A M Subbaiah, M., Kumar, B.V., Subramani, L., Wang, B., Li, Y.X., Sivaprakasam, P., Critton, D., Mulligan, D., Sandhu, B., Xie, C., Ramakrishnan, R., Nagar, J., Dudhgaonkar, S., Oderinde, M.S., Murtaza, A., Schieven, G.L., Mathur, A., Gavai, A.V., Vite, G., Gangwar, S.(2024) ACS Med Chem Lett 15: 181-188
- PubMed: 38352830 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.3c00455
- Primary Citation Related Structures:
8TTY, 8TTZ - PubMed Abstract:
We have designed and developed novel and selective TLR7 agonists that exhibited potent receptor activity in a cell-based reporter assay. In vitro , these agonists significantly induced secretion of cytokines IL-6, IL-1β, IL-10, TNFa, IFNa, and IP-10 in human and mouse whole blood. Pharmacokinetic and pharmacodynamic studies in mice showed a significant secretion of IFNα and TNFα cytokines. When combined with aPD1 in a CT-26 tumor model, the lead compound showed strong synergistic antitumor activity with complete tumor regression in 8/10 mice dosed using the intravenous route. Structure-activity relationship studies enabled by structure-based designs of TLR7 agonists are disclosed.
- Bristol-Myers Squibb Research & Development, 700 Bay Road, Redwood City, California 94063, United States.
Organizational Affiliation: