Human neutralizing antibodies target a conserved lateral patch on H7N9 hemagglutinin head.
Jia, M., Zhao, H., Morano, N.C., Lu, H., Lui, Y.M., Du, H., Becker, J.E., Yuen, K.Y., Ho, D.D., Kwong, P.D., Shapiro, L., To, K.K., Wu, X.(2024) Nat Commun 15: 4505-4505
- PubMed: 38802413 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-48758-4
- Primary Citation Related Structures:
8TNL, 8TOA - PubMed Abstract:
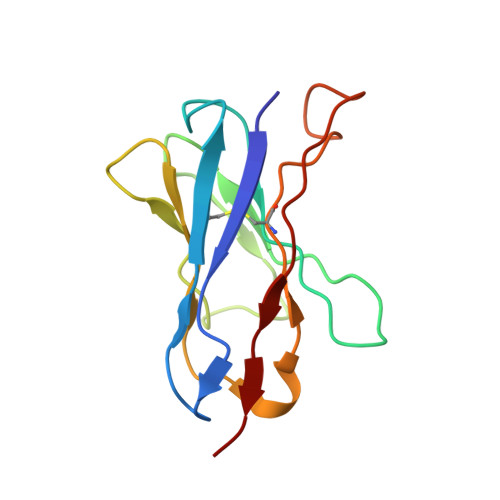
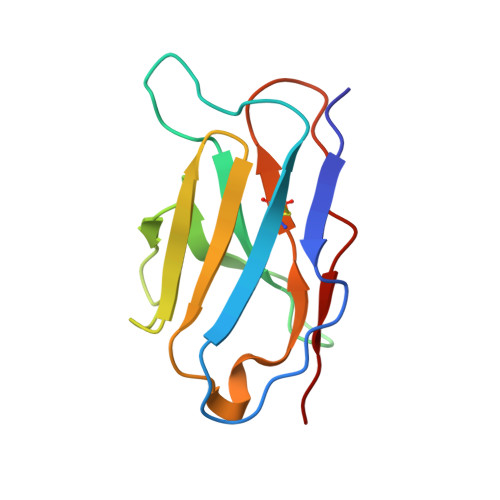
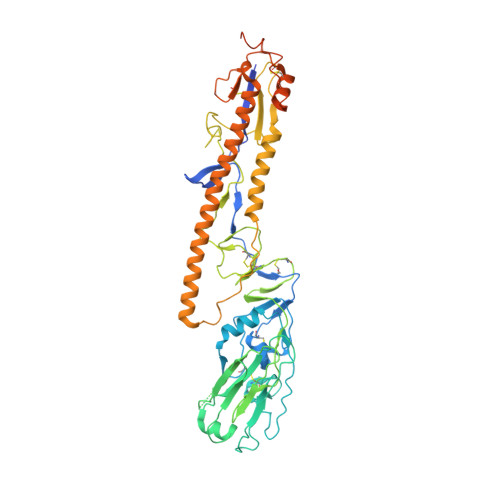
Avian influenza A virus H7N9 causes severe human infections with >30% fatality. Currently, there is no H7N9-specific prevention or treatment for humans. Here, from a 2013 H7N9 convalescent case in Hong Kong, we isolate four hemagglutinin (HA)-reactive monoclonal antibodies (mAbs), with three directed to the globular head domain (HA1) and one to the stalk domain (HA2). Two clonally related HA1-directed mAbs, H7.HK1 and H7.HK2, potently neutralize H7N9 and protect female mice from lethal H7N9/AH1 challenge. Cryo-EM structures reveal that H7.HK1 and H7.HK2 bind to a β14-centered surface and disrupt the 220-loop that makes hydrophobic contacts with sialic acid on an adjacent protomer, thereby blocking viral entry. Sequence analysis indicates the lateral patch targeted by H7.HK1 and H7.HK2 to be conserved among influenza subtypes. Both H7.HK1 and H7.HK2 retain HA1 binding and neutralization capacity to later H7N9 isolates from 2016-2017, consistent with structural data showing that the antigenic mutations during this timeframe occur at their epitope peripheries. The HA2-directed mAb H7.HK4 lacks neutralizing activity but when used in combination with H7.HK2 moderately augments female mouse protection. Overall, our data reveal antibodies to a conserved lateral HA1 supersite that confer neutralization, and when combined with a HA2-directed non-neutralizing mAb, augment protection.
- Aaron Diamond AIDS Research Center, Columbia University Vagelos College of Physicians and Surgeons, New York, NY, 10032, USA.
Organizational Affiliation: