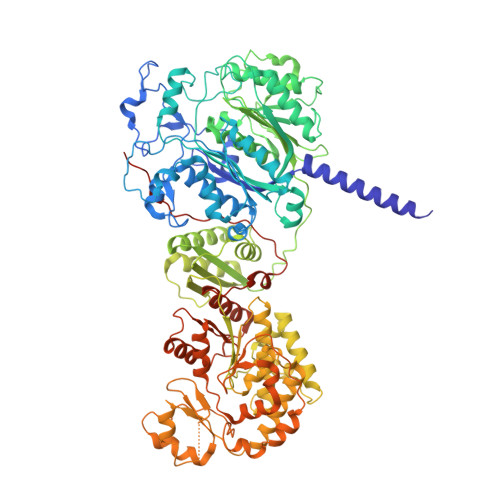
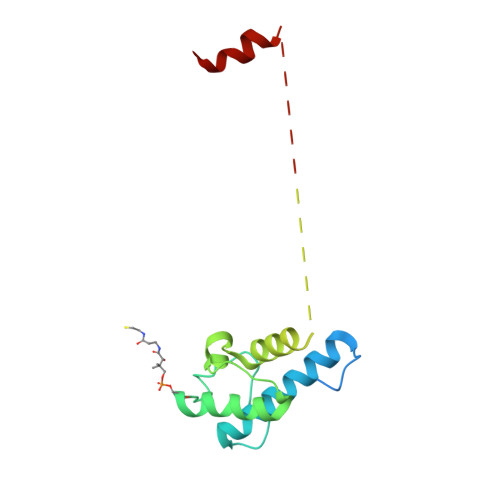
Structural basis for intermodular communication in assembly-line polyketide biosynthesis.
Cogan, D.P., Soohoo, A.M., Chen, M., Liu, Y., Brodsky, K.L., Khosla, C.(2025) Nat Chem Biol 21: 876-882
- PubMed: 39179672 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-024-01709-y
- Primary Citation Related Structures:
8TJN, 8TJO, 8TJP, 8TKO, 8TPW, 8TPX - PubMed Abstract:
Assembly-line polyketide synthases (PKSs) are modular multi-enzyme systems with considerable potential for genetic reprogramming. Understanding how they selectively transport biosynthetic intermediates along a defined sequence of active sites could be harnessed to rationally alter PKS product structures. To investigate functional interactions between PKS catalytic and substrate acyl carrier protein (ACP) domains, we employed a bifunctional reagent to crosslink transient domain-domain interfaces of a prototypical assembly line, the 6-deoxyerythronolide B synthase, and resolved their structures by single-particle cryogenic electron microscopy (cryo-EM). Together with statistical per-particle image analysis of cryo-EM data, we uncovered interactions between ketosynthase (KS) and ACP domains that discriminate between intra-modular and inter-modular communication while reinforcing the relevance of conformational asymmetry during the catalytic cycle. Our findings provide a foundation for the structure-based design of hybrid PKSs comprising biosynthetic modules from different naturally occurring assembly lines.
- Department of Chemistry, Stanford University, Stanford, CA, USA. dcogan@usc.edu.
Organizational Affiliation: