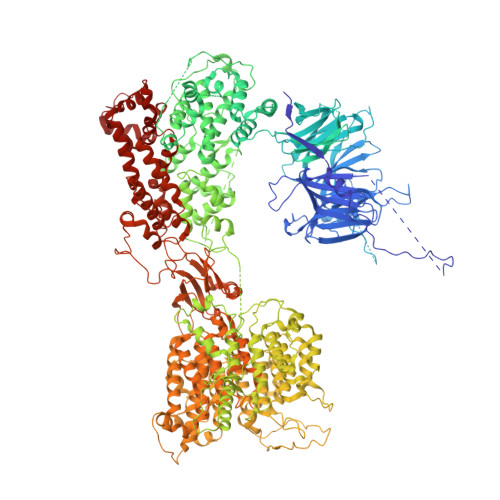
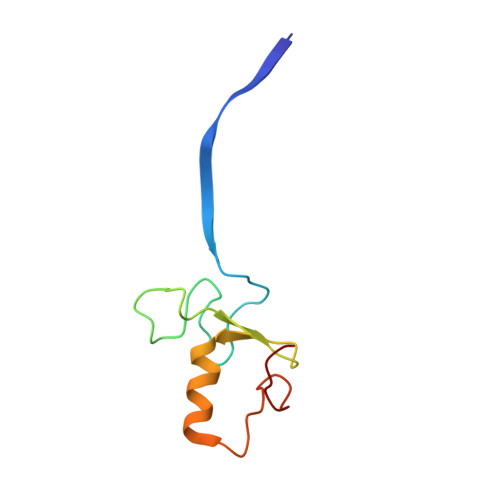
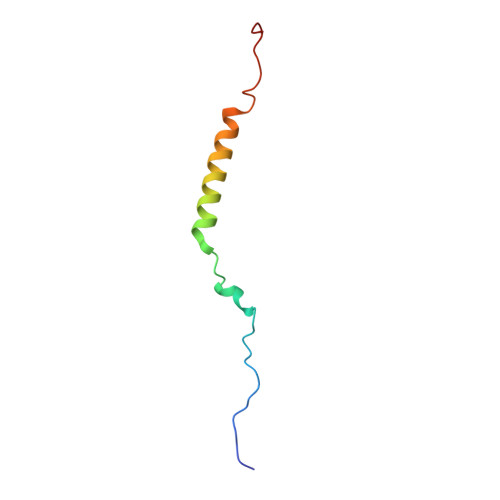
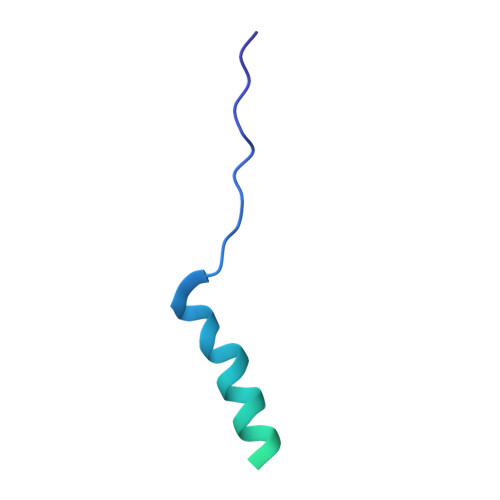
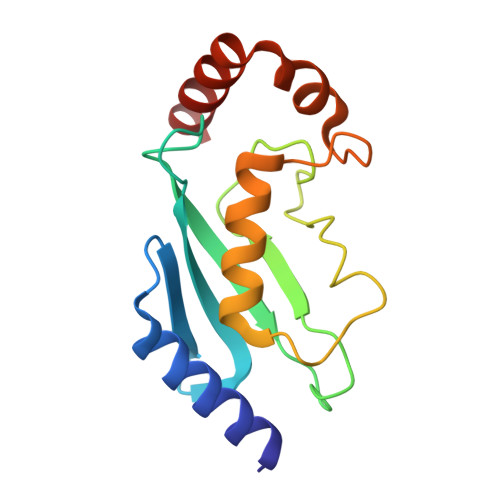
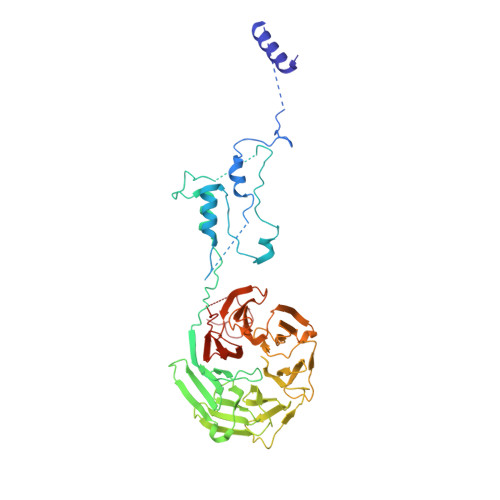
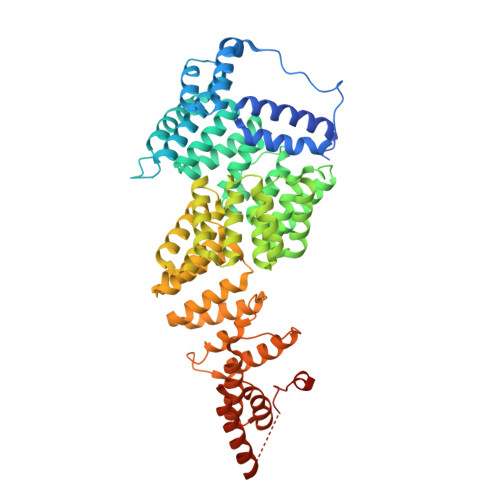
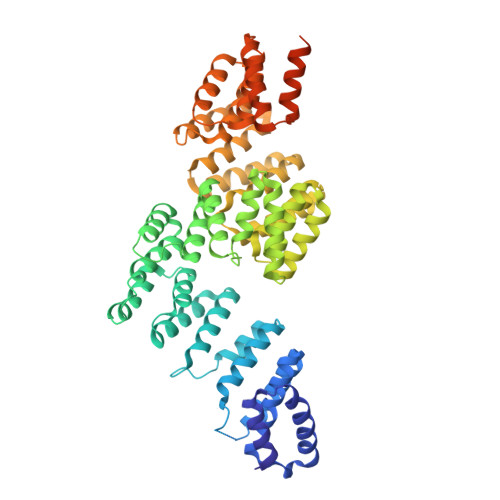
Time-resolved cryo-EM (TR-EM) analysis of substrate polyubiquitination by the RING E3 anaphase-promoting complex/cyclosome (APC/C).
Bodrug, T., Welsh, K.A., Bolhuis, D.L., Paulаkonis, E., Martinez-Chacin, R.C., Liu, B., Pinkin, N., Bonacci, T., Cui, L., Xu, P., Roscow, O., Amann, S.J., Grishkovskaya, I., Emanuele, M.J., Harrison, J.S., Steimel, J.P., Hahn, K.M., Zhang, W., Zhong, E.D., Haselbach, D., Brown, N.G.(2023) Nat Struct Mol Biol 30: 1663-1674
- PubMed: 37735619 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-023-01105-5
- Primary Citation Related Structures:
8TAR, 8TAU - PubMed Abstract:
Substrate polyubiquitination drives a myriad of cellular processes, including the cell cycle, apoptosis and immune responses. Polyubiquitination is highly dynamic, and obtaining mechanistic insight has thus far required artificially trapped structures to stabilize specific steps along the enzymatic process. So far, how any ubiquitin ligase builds a proteasomal degradation signal, which is canonically regarded as four or more ubiquitins, remains unclear. Here we present time-resolved cryogenic electron microscopy studies of the 1.2 MDa E3 ubiquitin ligase, known as the anaphase-promoting complex/cyclosome (APC/C), and its E2 co-enzymes (UBE2C/UBCH10 and UBE2S) during substrate polyubiquitination. Using cryoDRGN (Deep Reconstructing Generative Networks), a neural network-based approach, we reconstruct the conformational changes undergone by the human APC/C during polyubiquitination, directly visualize an active E3-E2 pair modifying its substrate, and identify unexpected interactions between multiple ubiquitins with parts of the APC/C machinery, including its coactivator CDH1. Together, we demonstrate how modification of substrates with nascent ubiquitin chains helps to potentiate processive substrate polyubiquitination, allowing us to model how a ubiquitin ligase builds a proteasomal degradation signal.
- Department of Biochemistry and Biophysics and Lineberger Comprehensive Cancer Center, University of North Carolina, Chapel Hill, NC, USA.
Organizational Affiliation: