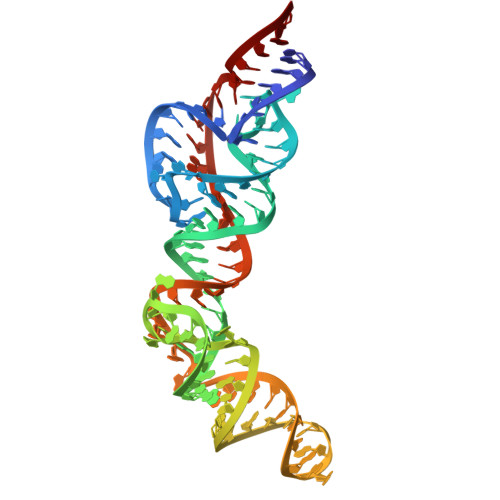
Structural investigation of an RNA device that regulates PD-1 expression in mammalian cells.
Stagno, J.R., Deme, J.C., Dwivedi, V., Lee, Y.T., Lee, H.K., Yu, P., Chen, S.Y., Fan, L., Degenhardt, M.F.S., Chari, R., Young, H.A., Lea, S.M., Wang, Y.X.(2025) Nucleic Acids Res 53
- PubMed: 40071935 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaf156
- Primary Citation Related Structures:
8SYK, 8T5O, 8TKJ, 8TKK - PubMed Abstract:
Synthetic RNA devices are engineered to control gene expression and offer great potential in both biotechnology and clinical applications. Here, we present multidisciplinary structural and biochemical data for a tetracycline (Tc)-responsive RNA device (D43) in both ligand-free and bound states, providing a structure-dynamical basis for signal transmission. Activation of self-cleavage is achieved via ligand-induced conformational and dynamical changes that stabilize the elongated bridging helix harboring the communication module, which drives proper coordination of the catalytic residues. We then show the utility of CRISPR-integrated D43 in EL4 lymphocytes to regulate programmed cell death protein 1 (PD-1), a key receptor of immune checkpoints. Treatment of these cells with Tc showed a dose-dependent reduction in PD-1 by immunostaining and a decrease in messenger RNA levels by quantitative PCR as compared with wild type. PD-1 expression was recoverable upon removal of Tc. These results provide mechanistic insight into RNA devices with potential for cancer immunotherapy or other applications.
- Protein-Nucleic Acid Interaction Section, Center for Structural Biology, Center for Cancer Research, National Cancer Institute, Frederick, MD, 21702, United States.
Organizational Affiliation: