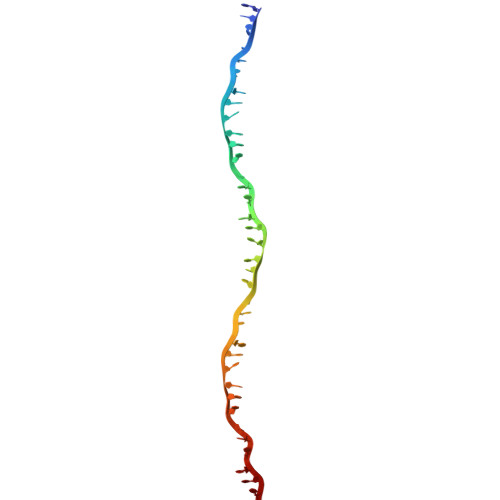
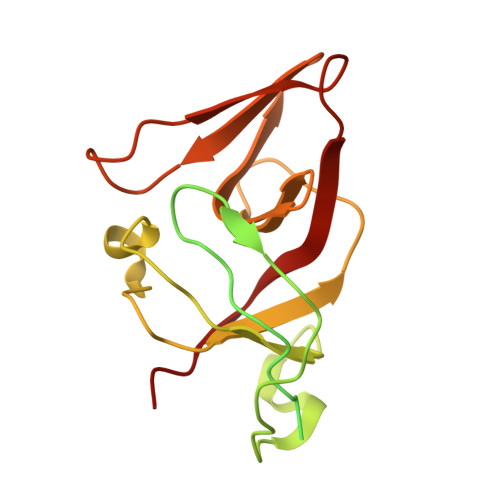
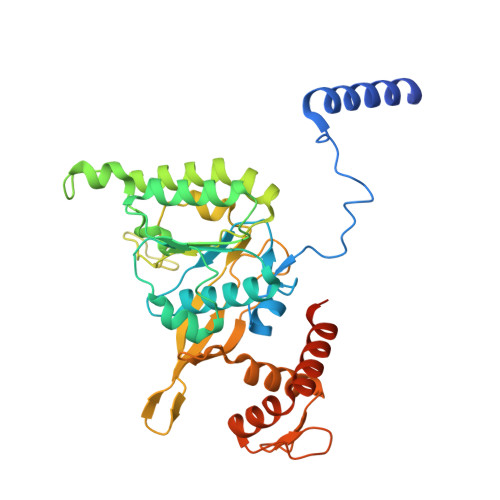
Snapshots of Pseudomonas aeruginosa SOS response reveal structural requisites for LexA autoproteolysis.
Vascon, F., De Felice, S., Gasparotto, M., Huber, S.T., Catalano, C., Chinellato, M., Mezzetti, R., Grinzato, A., Filippini, F., Maso, L., Jakobi, A.J., Cendron, L.(2025) iScience 28: 111726-111726
- PubMed: 39898034 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2024.111726
- Primary Citation Related Structures:
8B0V, 8S70, 8S7G - PubMed Abstract:
Antimicrobial resistance poses a severe threat to human health and Pseudomonas aeruginosa stands out among the pathogens responsible for this emergency. The SOS response to DNA damage is crucial in bacterial evolution, influencing resistance development and adaptability in challenging environments, especially under antibiotic exposure. Recombinase A (RecA) and the transcriptional repressor LexA are the key players that orchestrate this process, determining either the silencing or the active transcription of the genes under their control. By integrating state-of-the-art structural approaches with in vitro binding and functional assays, we elucidated the molecular events activating the SOS response in P. aeruginosa , focusing on the RecA-LexA interaction. Our findings identify the conserved determinants and strength of the interactions that allow RecA to trigger LexA autocleavage and inactivation. These results provide the groundwork for designing novel antimicrobial strategies and exploring the potential translation of Escherichia coli -derived approaches, to address the implications of P. aeruginosa infections.
- Department of Biology, University of Padua, Via Ugo Bassi 58/b, 35131 Padova, Italy.
Organizational Affiliation: