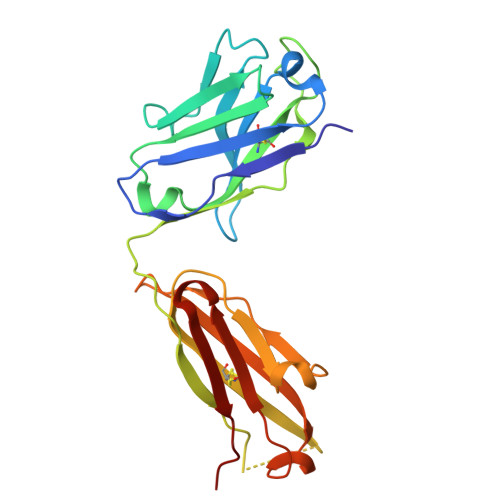
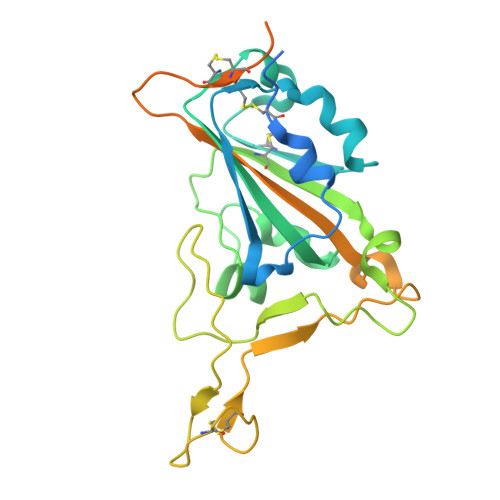
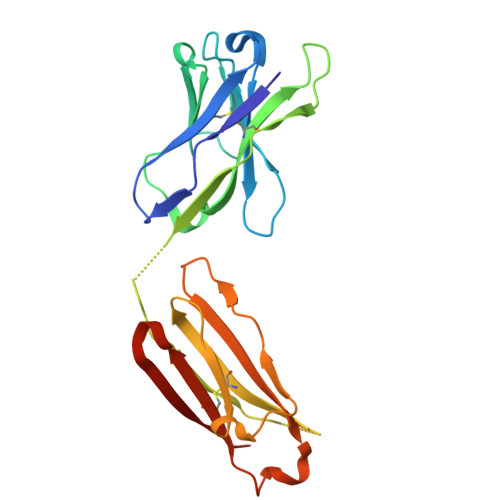
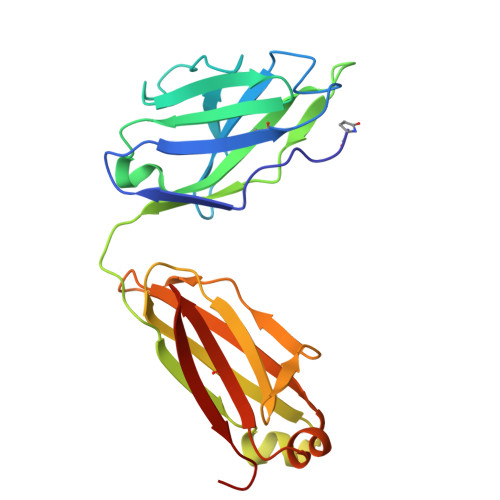
A potent pan-sarbecovirus neutralizing antibody resilient to epitope diversification.
Rosen, L.E., Tortorici, M.A., De Marco, A., Pinto, D., Foreman, W.B., Taylor, A.L., Park, Y.J., Bohan, D., Rietz, T., Errico, J.M., Hauser, K., Dang, H.V., Chartron, J.W., Giurdanella, M., Cusumano, G., Saliba, C., Zatta, F., Sprouse, K.R., Addetia, A., Zepeda, S.K., Brown, J., Lee, J., Dellota Jr., E., Rajesh, A., Noack, J., Tao, Q., DaCosta, Y., Tsu, B., Acosta, R., Subramanian, S., de Melo, G.D., Kergoat, L., Zhang, I., Liu, Z., Guarino, B., Schmid, M.A., Schnell, G., Miller, J.L., Lempp, F.A., Czudnochowski, N., Cameroni, E., Whelan, S.P.J., Bourhy, H., Purcell, L.A., Benigni, F., di Iulio, J., Pizzuto, M.S., Lanzavecchia, A., Telenti, A., Snell, G., Corti, D., Veesler, D., Starr, T.N.(2024) Cell 187: 7196
- PubMed: 39383863 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2024.09.026
- Primary Citation Related Structures:
8S6M, 9ASD, 9ATM, 9AU1, 9AU2 - PubMed Abstract:
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) evolution has resulted in viral escape from clinically authorized monoclonal antibodies (mAbs), creating a need for mAbs that are resilient to epitope diversification. Broadly neutralizing coronavirus mAbs that are sufficiently potent for clinical development and retain activity despite viral evolution remain elusive. We identified a human mAb, designated VIR-7229, which targets the viral receptor-binding motif (RBM) with unprecedented cross-reactivity to all sarbecovirus clades, including non-ACE2-utilizing bat sarbecoviruses, while potently neutralizing SARS-CoV-2 variants since 2019, including the recent EG.5, BA.2.86, and JN.1. VIR-7229 tolerates extraordinary epitope variability, partly attributed to its high binding affinity, receptor molecular mimicry, and interactions with RBM backbone atoms. Consequently, VIR-7229 features a high barrier for selection of escape mutants, which are rare and associated with reduced viral fitness, underscoring its potential to be resilient to future viral evolution. VIR-7229 is a strong candidate to become a next-generation medicine.
- Vir Biotechnology, San Francisco, CA 94158, USA.
Organizational Affiliation: