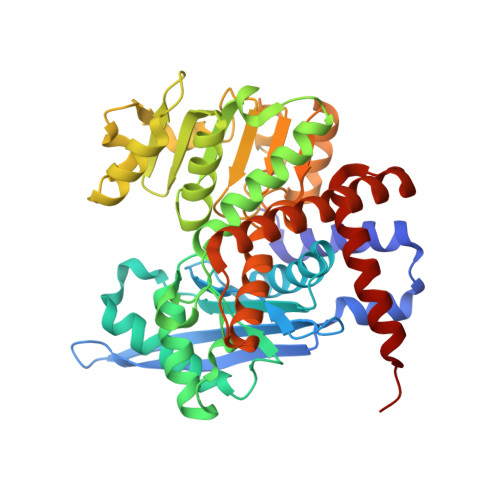
Legume-type glutamate dehydrogenase: Structure, activity, and inhibition studies.
Grzechowiak, M., Sliwiak, J., Link, A., Ruszkowski, M.(2024) Int J Biol Macromol 278: 134648-134648
- PubMed: 39142482 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2024.134648
- Primary Citation Related Structures:
8S38, 8S39, 8S3A, 8S3B, 8S3C, 8S3D - PubMed Abstract:
Glutamate dehydrogenases (GDHs) are key enzymes at the crossroads of N and C metabolism in plants. Legumes, whose N metabolism is particularly intricate, possess a unique type of GDH. This study presents an analysis of a legume-type GDH (isoform 2) from Medicago truncatula (MtGDH2). We measured MtGDH2 activity in both the Glu → 2-oxoglutarate (2OG) and 2OG → Glu reaction directions and obtained kinetic parameters for Glu, 2OG, NAD + , and NADH. Inhibition assays revealed that compounds possessing di- or tricarboxylates act as inhibitors of plant GDHs. Interestingly, 2,6-pyridinedicarboxylate (PYR) weakly inhibits MtGDH2 compared to Arabidopsis thaliana homologs. Furthermore, we explored tetrazole derivatives to discover 3-(1H-tetrazol-5-yl)benzoic acid (TBA) as an MtGDH2 inhibitor. The kinetic experiments are supported by six crystal structures, solved as: (i) unliganded enzyme, (ii) trapping the reaction intermediate 2-amino-2-hydroxyglutarate and NAD + , and also complexed with NAD + and inhibitors such as (iii) citrate, (iv) PYR, (v) isophthalate, and (vi) TBA. The complex with TBA revealed a new mode of action that, in contrast to other inhibitors, prevents domain closure. This discovery points to TBA as a starting point for the development of novel GDH inhibitors to study the functions of GDH in plants and potentially boost biomass production.
- Department of Structural Biology of Eukaryotes, Institute of Bioorganic Chemistry, Polish Academy of Sciences, Poznan 61-704, Poland.
Organizational Affiliation: