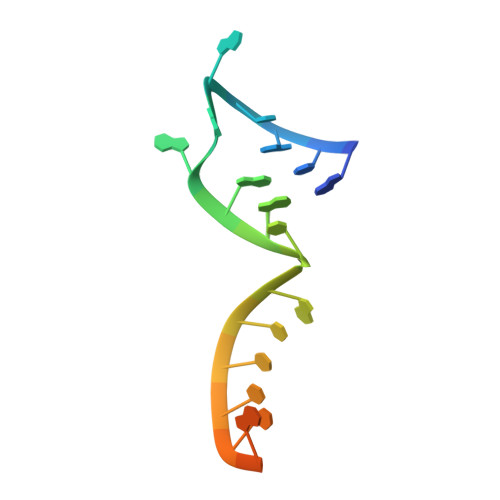
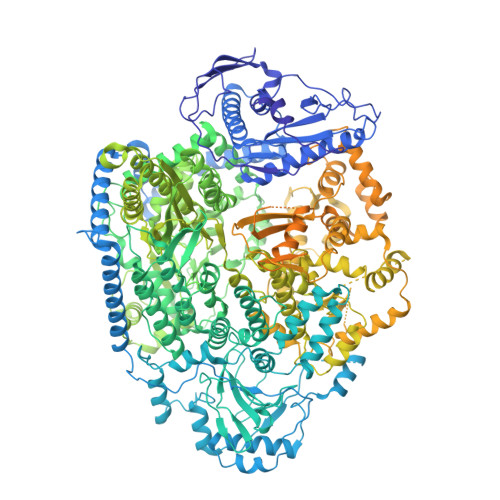
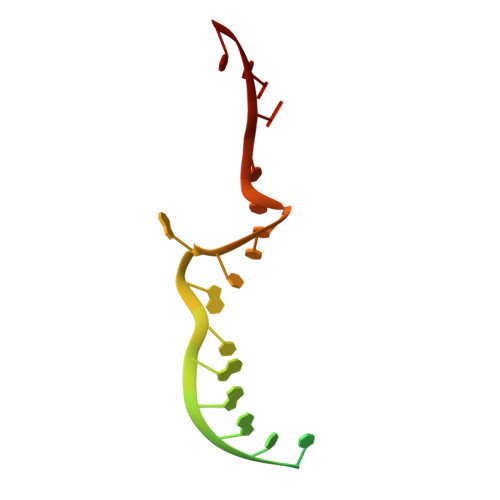
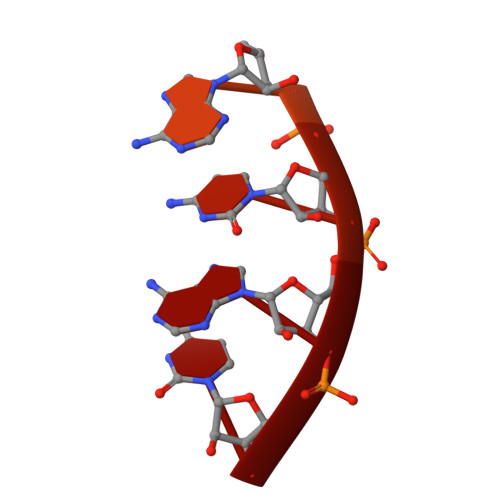
Structural snapshots of phenuivirus cap-snatching and transcription.
Williams, H.M., Thorkelsson, S.R., Vogel, D., Busch, C., Milewski, M., Cusack, S., Grunewald, K., Quemin, E.R.J., Rosenthal, M.(2024) Nucleic Acids Res 52: 6049-6065
- PubMed: 38709882 Search on PubMed
- DOI: https://doi.org/10.1093/nar/gkae330
- Primary Citation Related Structures:
8R6U, 8R6W, 8R6Y - PubMed Abstract:
Severe fever with thrombocytopenia syndrome virus (SFTSV) is a human pathogen that is now endemic to several East Asian countries. The viral large (L) protein catalyzes viral transcription by stealing host mRNA caps via a process known as cap-snatching. Here, we establish an in vitro cap-snatching assay and present three high-quality electron cryo-microscopy (cryo-EM) structures of the SFTSV L protein in biologically relevant, transcription-specific states. In a priming-state structure, we show capped RNA bound to the L protein cap-binding domain (CBD). The L protein conformation in this priming structure is significantly different from published replication-state structures, in particular the N- and C-terminal domains. The capped-RNA is positioned in a way that it can feed directly into the RNA-dependent RNA polymerase (RdRp) ready for elongation. We also captured the L protein in an early-elongation state following primer-incorporation demonstrating that this priming conformation is retained at least in the very early stages of primer extension. This structural data is complemented by in vitro biochemical and cell-based assays. Together, these insights further our mechanistic understanding of how SFTSV and other bunyaviruses incorporate stolen host mRNA fragments into their viral transcripts thereby allowing the virus to hijack host cell translation machinery.
- Bernhard Nocht Institute for Tropical Medicine (BNITM), Hamburg, Germany.
Organizational Affiliation: