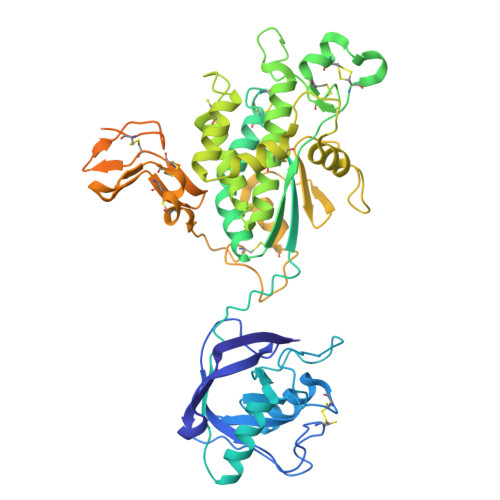
The crystal structure of Vacuolar Sorting Receptor 1 (VSR1) lumenal domain reveals a stable domain-swapped trimer and a potential new cargo binding site
Borges, R.J., Eldahshoury, M.K., Nettleship, J., Bird, L., An, J., Levada, A.J.L., Shah, N., Thomson, M., Fontes, M.R.M., Owens, R., Denecke, J., Postis, V., Uson, I., Goldman, A., De Marcos Lousa, C.To be published.