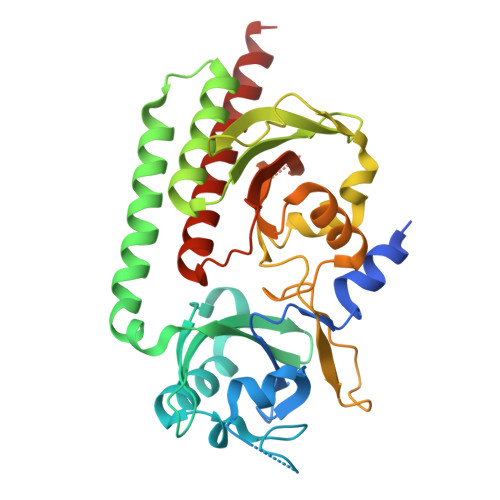
Pr and Pfr structures of plant phytochrome A.
Nagano, S., von Stetten, D., Guan, K., Chen, P.Y., Song, C., Barends, T., Weiss, M.S., Feiler, C.G., Dorner, K., de Diego Martinez, I., Schubert, R., Bielecki, J., Brings, L., Han, H., Kharitonov, K., Kim, C., Kloos, M., Koliyadu, J.C.P., Koua, F.H.M., Round, E., Sarma, A., Sato, T., Schmidt, C., Valerio, J., Wrona, A., Schulz, J., de Wijn, R., Letrun, R., Bean, R., Mancuso, A., Heyne, K., Hughes, J.(2025) Nat Commun 16: 5319-5319
- PubMed: 40544164 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-60738-w
- Primary Citation Related Structures:
8R44, 8R45, 9ER4, 9F4I, 9QZT - PubMed Abstract:
Phytochromes are biliprotein photoreceptors widespread amongst microorganisms and ubiquitous in plants where they control developmental processes as diverse as germination, stem elongation and floral induction through the photoconversion of inactive Pr to the Pfr signalling state. Here we report crystal structures of the chromophore-binding module of soybean phytochrome A, including ~2.2 Å XFEL structures of Pr and Pfr at ambient temperature and high resolution cryogenic structures of Pr. In the Pfr structure, the chromophore is exposed to the medium, the D-ring remaining α-facial following the likely clockwise photoflip. The chromophore shifts within its pocket, while its propionate side chains, their partners as well as three neighbouring tyrosines shift radically. Helices near the chromophore show substantial shifts that might represent components of the light signal. These changes reflect those in bacteriophytochromes despite their quite different signalling mechanisms, implying that fundamental aspects of phytochrome photoactivation have been repurposed for photoregulation in the eukaryotic plant.
- Institute for Plant Physiology, Justus Liebig University, Giessen, Germany.
Organizational Affiliation: