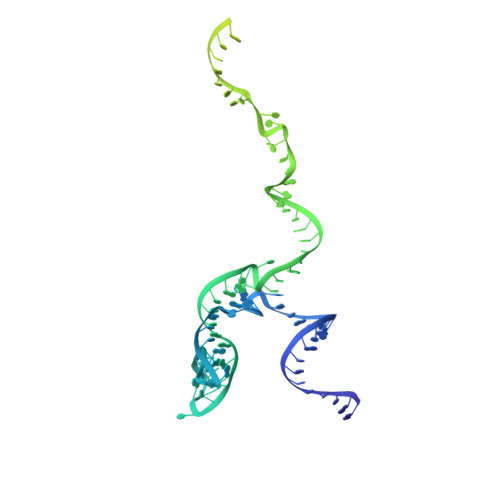
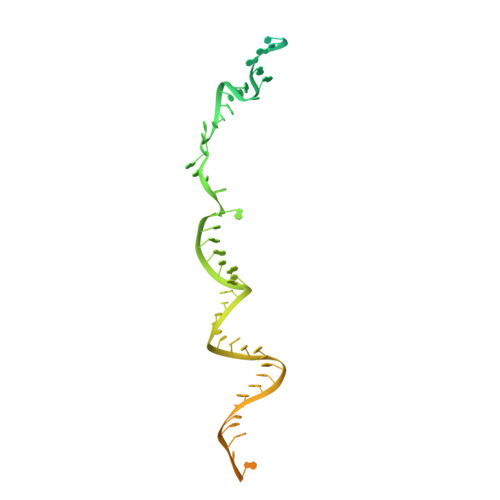
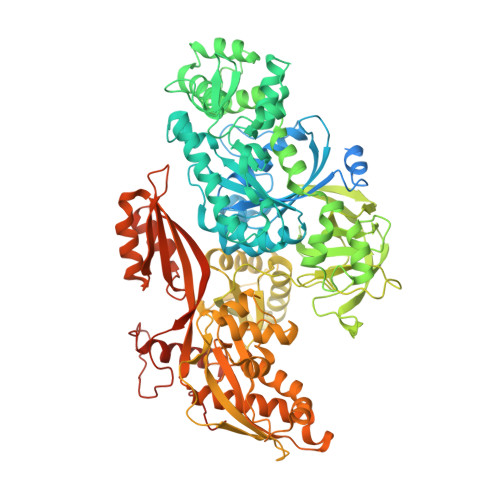
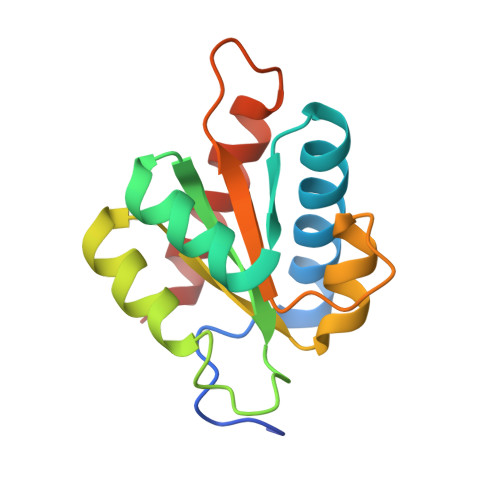
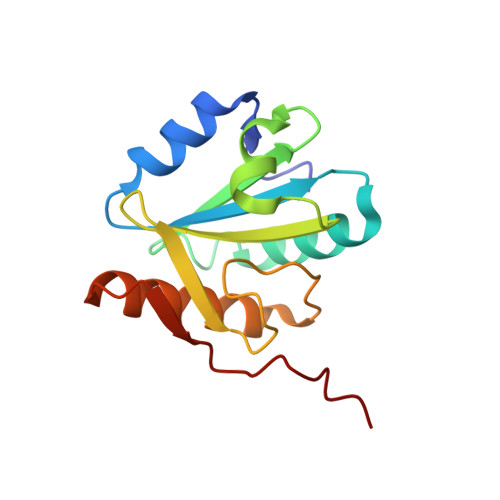
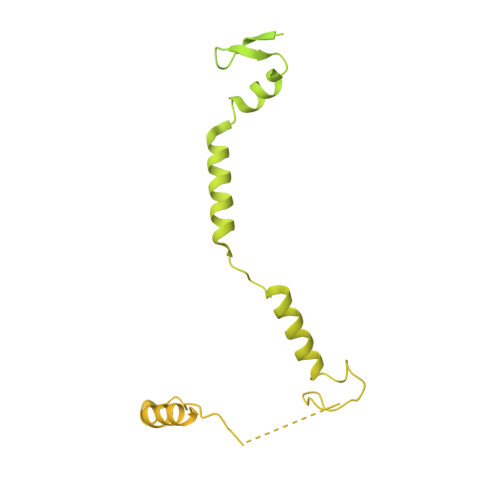
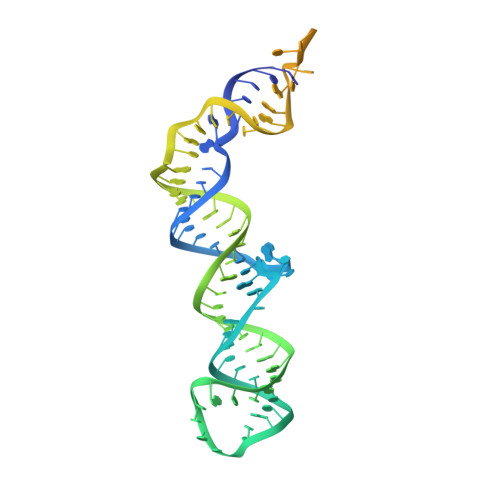
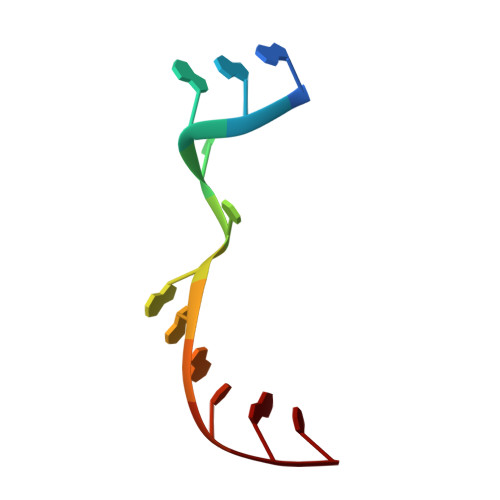Structural insights into the cross-exon to cross-intron spliceosome switch.
Zhang, Z., Kumar, V., Dybkov, O., Will, C.L., Zhong, J., Ludwig, S.E.J., Urlaub, H., Kastner, B., Stark, H., Luhrmann, R.(2024) Nature 630: 1012-1019
- PubMed: 38778104 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-024-07458-1
- Primary Citation Related Structures:
8QOZ, 8QP8, 8QP9, 8QPA, 8QPB, 8QPE, 8QPK, 8QXD, 8QZS, 8R08, 8R09, 8R0A, 8R0B, 8RM5 - PubMed Abstract:
Early spliceosome assembly can occur through an intron-defined pathway, whereby U1 and U2 small nuclear ribonucleoprotein particles (snRNPs) assemble across the intron 1 . Alternatively, it can occur through an exon-defined pathway 2-5 , whereby U2 binds the branch site located upstream of the defined exon and U1 snRNP interacts with the 5' splice site located directly downstream of it. The U4/U6.U5 tri-snRNP subsequently binds to produce a cross-intron (CI) or cross-exon (CE) pre-B complex, which is then converted to the spliceosomal B complex 6,7 . Exon definition promotes the splicing of upstream introns 2,8,9 and plays a key part in alternative splicing regulation 10-16 . However, the three-dimensional structure of exon-defined spliceosomal complexes and the molecular mechanism of the conversion from a CE-organized to a CI-organized spliceosome, a pre-requisite for splicing catalysis, remain poorly understood. Here cryo-electron microscopy analyses of human CE pre-B complex and B-like complexes reveal extensive structural similarities with their CI counterparts. The results indicate that the CE and CI spliceosome assembly pathways converge already at the pre-B stage. Add-back experiments using purified CE pre-B complexes, coupled with cryo-electron microscopy, elucidate the order of the extensive remodelling events that accompany the formation of B complexes and B-like complexes. The molecular triggers and roles of B-specific proteins in these rearrangements are also identified. We show that CE pre-B complexes can productively bind in trans to a U1 snRNP-bound 5' splice site. Together, our studies provide new mechanistic insights into the CE to CI switch during spliceosome assembly and its effect on pre-mRNA splice site pairing at this stage.
- Department of Structural Dynamics, Max-Planck-Institute for Multidisciplinary Sciences, Göttingen, Germany.
Organizational Affiliation: